- Home
- AD Genes
- AD risk factors
- Bipolar disorder
- BP Genes
- BP risk factors
- Schizophrenia
- SZ Genes
- SZ Risk factors
- Autism Risk factors
- Parkinson's disease risk factors
- Cystic Fibrosis
- Herpes simplex database
- Helicobacter pylori / host interactome
- T.Gondii/host interactome
- Jobs in genetics and bioinformatics
- Pathogens
- Pathways
- Useful links
- Subscribe
to Site Updates by Email
 Subscribe
in a reader
Subscribe
in a reader
KEGG pathway analysis of over 400 Alzheimer's disease susceptibility genes Other KEGG analyses in other diseases
- Alzheimer's disease: a pathogenetic autoimmune disorder caused by herpes simplex in a gene-dependent manner. Int.J.Alz.Dis. 2010
- APP, APOE, complement receptor 1, clusterin and PICALM and their involvement in the herpes simplex life cycle. Neurosci Lett. 2010 ;483(2):96-100.
- The Fox and the Rabbits�Environmental Variables and Population Genetics (1) Replication Problems in Association Studies and the Untapped Power of GWAS (2) Vitamin A Deficiency, Herpes Simplex Reactivation and Other Causes of Alzheimer's Disease ISRN Neurology 2011
- APP, Gamma Secretase, APOE, CLU, CR1, PICALM,ABCA7, BIN1, CD2AP, CD33, EPHA1, and MS4A2, and Their Relationships with Herpes Simplex, C. Pneumoniae, Other Suspect Pathogens, and the Immune System. Int JAlzheimers Dis. 2011;2011:501862.
Alzheimer�s disease plaques and tangles: Cemeteries of a Pyrrhic victory of the immune defence network against herpes simplex infection at the expense of complement and inflammation�mediated neuronal destruction.
Summary
Introduction
Alzheimer�s disease is a devastating degenerative disorder characterised by extensive neuronal loss particularly of the cholinergic system (Sims, Bowen et al. 1980) , severance of the afferent and efferent hippocampal connections (Van Hoesen and Hyman ,1990) , loss of corticocortical glutamatergic association fibres (Young ,1987) and massive cerebral shrinkage (Thompson, Hayashi et al. 2003) . It is characterised pathologically by the presence of amyloid containing plaques (Nikaido, Austin et al. 1970) and neurofibrillary tangles (Ishino and Otsuki ,1975) containing the hyperphosphorylated microtubule protein, tau (Kosik ,1990) . Plaques and tangles are thought by many to be the factors leading to the neuronal devastation in this disease (Perl ,2010). An understanding of their genesis is a key goal in Alzheimer�s disease research. These features, including beta-amyloid deposition,� tau phosphorylation (Itzhaki, Cosby et al. 2008) (Piacentini, Civitelli et al. 2010;Wozniak, Frost et al. 2009) , entorhinal and hippocampal cell loss and cerebral shrinkage (Armien, Hu et al. 2009) can all be induced by herpes simplex viral infection in mice.
Herpes simplex can enter the brain via neural routes, typically via the trigeminal nerve, from whence the virus targets the limbic system (Becker ,1995;Damasio and Van Hoesen ,1985) , but can also enter via the bloodstream. Infection via this route is favoured by the possession of the APOE4 variant in mice and also by gender, with female mice showing a greater degree of cerebral infection (Burgos, Ramirez et al. 2005;Burgos, Ramirez et al. 2006;Burgos, Ramirez et al. 2003;Burgos, Ramirez et al. 2002) �. The virus binds to all classes of lipoprotein (VLDL, LDL and HDL) and the viral glycoprotein B binds to APOA1 and APOE containing lipoproteins (Huemer, Menzel et al. 1988) . The virus also binds to complement receptor 1 (CR1) on erythrocytes (Powers, Buster et al. 1995) . As CR1 is expressed in phagocytic Kolmer cells in the choroid plexus (Singhrao, Neal et al. 1999) , this receptor, a key player in Alzheimer�s disease genetics (Lambert, Heath et al. 2009) , �could provide a means of viral cerebral entry.
There is evidence for herpes simplex infection (HSV-1) as a risk factor in Alzheimer�s disease, acting in synergy with possession of the apolipoprotein E APOE4 allele (Itzhaki, Lin et al. 1997) . A recent study has also shown that anti-HSV-1 immunoglobulin M seropositivity, a marker of primary viral infection or reactivation, in a cohort of healthy patients, was significantly associated with the subsequent development of Alzheimer�s disease. Anti-HSV-1 IgG, a marker of lifelong infection showed no association with subsequent Alzheimer�s disease development (Letenneur, Peres et al. 2008) .� Herpes simplex viral DNA is also found in the beta-amyloid containing plaques that characterise Alzheimer�s disease (Wozniak, Mee et al. 2009) ,
�During the course of viral infection, the herpes virus engages with a number of human proteins which it uses to gain cellular entry, and to traffic from the cell membrane to the nucleus and back, via endosomal and other compartments. It also uses the host�s transcriptional machinery to replicate, and binds to proteins that control immune surveillance or apoptosis. Numerous evasion strategies related to these interactions regulate the survival and propagation of the virus. Concomitantly, host proteins attempt to destroy the virus, and the eventual outcome of this host/pathogen war determines the success of the virus, and the degree of damage it may cause. Paradoxically, successful viral elimination by the host�s immune system could well lead to greater cellular damage, mediated via immune and inflammatory targeting of the cells containing the virus.
A literature survey of herpes simplex binding proteins and of the proteins identified in Alzheimer�s disease plaques or neurofibrillary tangles (NFT�s) identified a large number of HSV-1 related proteins in these structures. These proteins relate to all stages of the viral life cycle .Both structures also contain a number of immune related proteins, suggesting that they may represent the record of a host/viral battle, whose outcome is viral elimination at the expense of massive neuronal destruction.
�
Methods
HSV-1 viral binding chemicals and proteins were identified by literature survey and are stocked and referenced in a database at http://www.polygenicpathways.co.uk/herpeshost.html
�which serves as the reference source for the viral/host interactions described below. At the time of writing, the database contains 341 direct interactions between the virus and host proteins or chemicals and a number of other effects such as mitochondrial gene deletion. Indirect effects, such as those on gene or protein expression, are not included. The constituents of amyloid containing plaques or neurofibrillary tangles (NFT�s) in Alzheimer�s disease were trawled by literature survey and a large number were identified from a proteomics study of laser-dissected plaques, which identified 488 proteins within these structures (Liao, Cheng et al. 2004) and from a similar study of� NFT�s which reported 79 tangle related proteins (Wang, Woltjer et al. 2005). These datasets were used to cross reference the protein accession numbers of HSV-1 interacting proteins.� Genes implicated in Alzheimer�s disease are referenced at http://www.polygenicpathways.co.uk/alzpolys.html. Gene symbols recognised by the Human Gene Nomenclature committee (HUGO) are used throughout the text and figures, and full names are given in Tables 3 -8.
�
Results
Statistical analysis
and the types of viral interacting protein found in plaques and tangles
For the two proteomics studies, HSV-1 binding proteins were highly and significantly enriched in both the plaque (10.7 fold: p=2.5229E-125) and tangle (14.4 fold:� p=4.47466E-39) datasets. Conversely, plaque and tangle components were highly and significantly enriched in the HSV-1 binding dataset (Plaque proteins = 7.4 fold: p=5.19975E-77: Tangle proteins = 61.6 fold: P= 9.5142E-39) (Table 1). Viral associated proteins were also second of the functional classes observed in plaques and first if mitochondrial proteins are included (mitochondria cluster around the virus at intracellular infection sites (Table 1, see below).
64 viral associated proteins were present in the plaque proteome and 14 were present in the tangle proteome (19% and 4% respectively of the known HSV-1 binding proteins). Further proteins in plaques and tangles were identified by literature survey .111 viral associated proteins were identified as amyloid plaque components, and 68 as tangle components. In total 132 exact protein/chemical matches (39% of the known viral associated proteins) were identified in either compartment.
A summary of the types of protein found in plaques or tangles is provided in Table 2 and references for their association with either of these structures in Tables 3 to 8 which are organised by functional category. These categories are illustrated in Figs 1-4.
There was considerable overlap between the two compartments as shown in Table 2, although the profiling of the proteins in relation to function was distinct in the two compartments (Figs 3-4). However a number of proteins were plaque (67) or tangle (23) specific. These are highlighted in Table 2, and illustrated in Figs 1-4. Moreover, the viral interacting proteins found in plaques and tangles are clustered in distinct functional classes sequentially related to different phases of the viral life cycle (Table 2, Figs 1-4). For example, almost all of the known carriers and primary receptors for the herpes simplex virus (100 and 63% respectively) are present in plaques or tangles (Table 2) and a high proportion of host viral associated proteins related to herpes simplex endocytosis (91%), intervesicular transport (88%), anterograde and retrograde transport (85 and 94%) and host virion components (71%) are also present in these structures. Host nuclear proteins, or those binding to the viral genome, or latency transcript, are much less represented (Fig 3, 4) (see below).
Viral associated
proteins in amyloid plaques (Fig 1, Fig 3)
Amyloid plaques contain numerous herpes simplex carriers, (defined as soluble molecules occupying the extracellular space), including APOE, as well as a large number of known viral receptors (Table 2,3, Fig 1). Two of these, syndecans 1 and 2, are substrates for the APP protease complex, gamma-secretase (Hemming, Elias et al. 2008).
Herpes simplex binds to heparan sulphate and chondroitin sulphate proteoglycans; although within this family only syndecans 1 and 2 have been specifically tested as entry receptors (Cheshenko, Liu et al. 2007) . Agrin, Aggrecan, Decorin, Glypican 1, Perlecan, syndecans 3 and 4, neurocan and Versican belong to these families and are all present in plaques or tangles. These might all be considered as potential HSV-1 receptors (Table 2, 3).as could a chondroitin sulphate proteoglycan form of APP, appican, which is expressed in astrocytes (Pangalos, Shioi et al. 1996) . Calculations with or without these receptors are provided in Tables 1 and 2.
The cellular uptake of the herpes virus protein VP22 is mediated via lipid raft-dependent endocytosis that depends on dynamins and ADP ribosylation factors. Dynamin 1 (DNM1) is expressed in both plaques and tangles and numerous ARF�s (virion components), which play a general role in membrane traffic (D'Souza-Schorey and Chavrier ,2006) are predominantly expressed in amyloid containing plaques (Table 4, Fig 1). The routing of viral glycoprotein D to the endosomal compartment is also mediated by mannose-6-phosphate receptors (IGF2R and M6PR) both of which also play a role in viral entry (Table 4, Fig 1). �(Brunetti, Dingwell et al. 1998) Clusterin is a ligand for M6PR (Lemansky, Brix et al. 1999) while PICALM (Phosphatidyl inositol binding clathrin assembly protein) overexpression reduces its endosomal localisation, suggesting blockade of its transport from the plasma membrane or the trans-Golgi network to endosomes (Tebar, Bohlander et al. 1999)(Fig 1) .Thus plaques contain many elements of the machinery delivering the virus to intracellular vesicular compartments. Two major Alzheimer�s disease susceptibility genes, clusterin and PICALM) are implicated in this machinery, while APOE is involved in serum viral transport.
Retrograde viral transport towards the nucleus (Table 4, Fig1) is mediated by dynein motors which carry the virus and walk it along the microtubule network. The dynein motor is composed of numerous dynein light, intermediate and heavy chains and three dynactins. The herpes virus binds to dynactin1 (p150/glued) and to dyneins DYNC1I1, DYNLL1, DYNLT1 and DYNLT3, although other components of the motor are likely to be involved in viral transport (Lyman and Enquist ,2009). The amyloid plaque contains the viral binding dynein DYNC1I1 and DYNLL1 (as well as DYNC1H1, and DYNLL2, which have not been shown to bind to the virus but which are generally involved in retrograde transport) (Fig 1).
The herpes simplex virion associates with host actins (ACTB, ACTG1), cofilin 1, which disassembles actin filaments,� profilin1, which regulates actin polymerisation (Bugyi and Carlier ,2010) and cysteine rich protein , CSRP1, �which binds to the actin cross-linking protein, actinin (Harper, Beckerle et al. 2000).� Actins also play a role in HSV-1 intracellular transport including within, and to and from the nucleus (Lyman and Enquist ,2009). Nuclear export of the virus is mediated by crm1 (exportin XPO1) �a protein that binds to the product of a major Alzheimer�s disease susceptibility gene, PICALM. This is discussed below (Fig 1).
The anterograde transport� (nucleus to cell surface) of the virus along microtubules involves APP (Satpute-Krishnan, DeGiorgis et al. 2003) (see below) and kinesin motors (Lyman and Enquist ,2009) (Table 4, Fig 1). The kinesin-1 (KIF1A or KIF5B) viral associated proteins were not found the proteomics study of amyloid plaques, although kinesin light chain KLC3 was present. Kinesins also play a role in APP transport, which is discussed below. Multiple forms (45) of kinesin exist, and their individual role in herpes viral or APP transport, or in plaque deposition remains to be assessed.
Numerous virion-incorporated RAB proteins (members of the Ras superfamily of monomeric G proteins) are amyloid plaque components (Table 5). These play multiple roles in intracellular traffic (endoplasmic reticulum, Golgi, endosome, lysosome and autophagosome). �Their distribution in these different compartments, based on recent reviews (Stenmark ,2009;Cardoso, Jordao et al. 2010), is illustrated in Fig 1. The trans-Golgi network protein TGOLN2 is also involved in protein transport within the trans-Golgi network, endosomes, and the cell membrane (McNamara, Grigston et al. 2004). Many viral associated proteins including RABS and the autophagy protein beclin 1 are present in these intervesicular compartments.
����������� Several annexins are both virion and plaque components (Table 5) and ANXA2 has also been isolated as a virion associate from HSV-1 infected cells (Padula, Sydnor et al. 2009). Annexins are involved in endo- and exocytosis while the growth cone associated protein, GAP43 and the synaptosomal protein, SNAP25, are involved in viral exocytosis (Fatimathas and Moss ,2010).
A recent proteomics study of 14-3-3 proteins delivered many binding partners. Prominent among these were a number of kinesins (KIF1B, KIF1C, KIF23, KLC2, KLC3, KLC4 and the viral binding KIF5B) and tubulins (TUBB, TUBB4 and TUBB4A), shared between YWHAE, YWHAG and YWHAZ. YWHAG also bound to dynein (DYNC1H1) (Jin, Smith et al. 2004). 14-3-3 proteins have many diverse roles but their relationship to transport systems is highlighted in Figs 2 and 3. Their positioning relates to the binding partners in the proteomics publication. �
In addition to these transport related proteins, amyloid plaques contain several viral binding protein chaperones (calnexin and heat shock proteins), as well as ubiquitin and the proteasomal unit PSMA2 and the translation initiation factor, EIF4A2. (Table 7, 8) .The chaperone/proteasome/ubiquitin machinery is hijacked by herpes simplex which moves these complexes into discrete nuclear foci probably involved in the quality control of viral proteins (Burch and Weller ,2004). The ubiquitin proteasome system, necessary for the elimination of abnormal proteins such as beta-amyloid and tau, is dysfunctional in Alzheimer�s disease (Paul ,2008), a situation that might well be induced by the effects of the virus on the chaperone/proteasome/ubiquitin machinery. A number of viral binding nuclear proteins (BRCA1, HNRNPK, APEX1, PARP1, XRCC6, and numerous Histones) are likely to be involved in viral DNA replication. Clusterin appears in this compartment as it binds to the Ku antigen and DNA repair protein (XRCC6) in nuclei, an interaction promoting apoptosis (Leskov, Klokov et al. 2003).
Plaques contain a number of viral binding coagulation factors. These are part of a cascade eventually leading to thrombin activation. Thrombin is involved in the proteolysis of APP, generating a c-terminal derived peptide that may be a precursor for beta-amyloid (Igarashi, Murai et al. 1992) (Fig 5).�����
Other classes of viral associated proteins included metabolic proteins (Glyceraldehyde 3-phosphate dehydrogenase, triosephosphate isomerase and nucleoside diphosphate kinase A). Several viral binding free radical related proteins, peroxiredoxin and thioredoxin are also plaque components. Immune and defence related viral associated proteins are well-represented plaque components ( Table 6) including complement C3, cyclophilin (PPIA) the viral DNA activated kinase EIF2AK2 , also known as protein kinase R, and its activator PRKRA,� immunoglobulin G , Interleukin 6, macrophage inhibitory factor and� HLA-antigens. The proteasome unit PSMA2 is also involved in the immune system as part of the immunoproteasome that processes MHC peptides (Wang and Maldonado ,2006). XRCC6 is included in this list as it doubles as a protein involved in somatic recombination, a process that generates multiple T cell receptors and immunoglobulins that recognise viruses and other pathogens (Gu, Jin et al. 1997).
The proteomics study of amyloid containing plaques reported a high proportion (53/488: 11%) of mitochondrial elements related to the Krebs cycle and oxidative phosphorylation (Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) a result that might reflect the clustering of� mitochondria around the herpes simplex virus following infection (Bello-Morales, Fedetz et al. 2005). �In addition many enzymes related to glycolysis (18) and pyruvate metabolism (8) were observed in amyloid plaques. HSV-1 infection is known to increase cerebral glucose consumption in infected brain areas (Saito and Price ,1984).
�The viral associated proteins found in amyloid plaques effectively, and sequentially, trace out the whole process of the viral life cycle, from carriage and entry, through transport to nuclear or endosomal compartments, nuclear DNA replication, nuclear egress, protein synthesis and packaging, anterograde transport and release. Along this route, the sequestration of many proteins by the virus is likely to promulgate its traffic through diverse neuronal compartments. In addition, such sequestration is likely to influence such diverse effects as APP traffic, neurotransmitter release, neurite outgrowth, neuronal survival, neuroprotection, inflammation, and astrogliosis � all key processes disrupted in the Alzheimer�s disease brain.�
Viral associated
proteins in neurofibrillary tangles (Fig 2, Fig 4)
The proteomics laser dissection study of NFT�s reported much fewer proteins (79) than the plaque study. 14 of which (18%) exactly match known viral associated proteins. HSV-1 also binds to tubulins, although only TUBA1B has been specifically reported. Several members of this family were NFT components in this study (TUBA4A, TUBB2A, TUBB3, TUBB4 and TUBB4Q). The virus also binds to Histones, 11 of which were present in this NFT proteomics study (Table 6). These are involved in DNA packaging. As in plaques, the chondroitin sulphate proteoglycan, Versican was present in NFT�s. If these proteins are included, 30/79 (38%) tangle components can be related to the virus (Table 1).
Other studies have shown that the viral binding proteins APOE and fibroblast growth factor ,FGF2, and the viral receptors Heparan sulphate, Chondroitin sulphate , FGFR1, IGF2R, syndecans SDC1, SDC2 as well as heparan sulphate and chondroitin sulphate proteoglycans are present in NFT�s, �as in plaques (Table 2, 3, Fig 2). Viral associated proteins related to endocytosis, retrograde, nuclear and anterograde transport, including the microtubule associated protein tau (MAPT), are also well represented in tangles (Table 2, 4, Fig 2, 4).
The viral binding kinases CDC2 and casein kinase CSNK2B �both phosphorylate tau (Singh, Grundke-Iqbal et al. 1994) while the viral binding protein, SET, is a phosphatase inhibitor that regulates tau dephosphorylation (Chohan, Khatoon et al. 2006). HSV-1 infection also increases glycogen synthase kinase GSK3B and protein kinase a (PRKRCA) expression and tau phosphorylation (Wozniak, Frost, and Itzhaki ,2009).
As with plaques, NFT�s contain nuclear proteins, heat shock proteins, translation initiation factors, 14-3-3 proteins, GAPDH , and peroxiredoxins. The binding of 14-3-3 proteins to tubulins and kinesins has been covered above. In addition YWHAZ, the predominant form in tangles, binds to tau (MAPT) and enhances its phosphorylation by cAMP dependent kinase (Hashiguchi, Sobue et al. 2000). Tangles also contain ubiquitin and proteasome components. As with plaques, tangles contain a number of immune/defence related proteins including complement C3, the membrane attack complex inhibitor CD59, cyclophilin, HLA antigens and the viral activated kinase EIF2AK2.
The virus uses the microtubule network for both anterograde and retrograde transport, and its association with a number of elements related to tau phosphorylation are likely to impact upon microtubule function.
Differences
in the functional profiles of viral-binding proteins in plaques and tangles
(Fig 3, Fig 4).
There is an interesting gradient in relation to the types of viral proteins that bind to the host proteins, which is similar in both amyloid plaques and tangles; glycoprotein > virion > capsid > tegument > viral genome for plaques, and glycoprotein > virion > tegument > viral genome > capsid for tangles, �roughly corresponding to the layering and sequential processing of the virus. Plaques contain a higher proportion of viral receptors and carriers, and of the viral associated proteins involved in endocytosis, intervesicular transport and exocytosis. They also specifically contain host proteins binding to the viral latency transcript (Fig 3, 4), suggesting that plaque components may be involved in viral reactivation.
Amyloid plaques are also enriched in the viral associated proteins involved in APP and beta-amyloid processing (Fig 3, 4), and contain 53 mitochondrial related proteins (Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) which may reflect mitochondrial clustering around the virus after infection (Bello-Morales, Fedetz, Alcina, Tabares, and Lopez-Guerrero ,2005). Tangles are relatively enriched in viral associated proteins related to tau phosphorylation, and anterograde and retrograde transport (Fig 4), as might be expected. Tangles also contain relatively higher proportions of viral-binding proteins involved in signalling, although these are also mostly related to tau phosphorylation.
In both cases, nuclear related viral binding host proteins (replication, repair, transcription, chromatin, or proteins binding to the viral single stranded DNA binding protein ICP8, (involved in viral genome replication and recombination) are relatively poorly represented. A speculative interpretation of the sequential ordering of the viral compartments is that viral replication, entailing host nuclear compartments, has not been very successful and that the structural components of the virus (glycoprotein, virion, tegument, and capsid) have been eliminated. In this respect, both plaques and tangles contain relatively high proportions of autophagy and immune related viral associated proteins (Table 7 , Fig 3, 4) (see below). Autophagy is the process of cellular self-digestion mediated by lysosomes, that also plays a more controlled role in viral destruction and in the destruction of aberrant proteins such as beta-amyloid. This system is disrupted in the Alzheimer�s disease brain(Kim, Lee et al. 2010) (Itzhaki, Cosby, and Wozniak ,2008).
�
Immune related proteins in amyloid plaques and neurofibrillary tangles (Figs 1- 5).
As well as the immune or defence-related herpes simplex binding proteins covered in this survey, a number of other immune-system related proteins are found in amyloid plaques or NFT�s. The cytokines, Interleukin 1 alpha, IL6 and tumour necrosis factor TNF have also all been localised within amyloid containing plaques (Veerhuis, Janssen et al. 1999). Acute phase proteins involved in inflammation, such as amyloid P, alpha-1 antichymotrypsin and C-reactive protein are also plaque components (Eikelenboom, van Exel et al. 2010) while Immunoglobulin G is located in the plaque corona (Eikelenboom and Stam ,1982). �
As well as The herpes simplex
binding protein Complement C3, a number of other complement related proteins
are found in amyloid plaques or NFT�s. Complement components Clq, C3d, and C4d
are present in plaques, dystrophic neurites and NFT�s (McGeer, Akiyama et al.
1989).
�The membrane attack complex (MAC: complements
complex C5b, C6, C7 C8 C9) is not
present in amyloid plaques,
but is observed in dystrophic neurites and tangle containing neurones (McGeer, Akiyama,
Itagaki, and McGeer ,1989) . The MAC complex has also
been detected in the neuronal cytoplasm in AD brains, associated with NFT�s and
lysosomes, a distribution that suggests endocytosis of membrane bound MAC and
its retrograde transport to the lysosome (Itagaki, Akiyama et al.
1994). The MAC
complex is a channel that is inserted into pathogen membranes, or into host
cell membranes, causing death by osmotic-related lysis. The complement inhibitors,
complement receptor 1 (CR1), CD59,
decay accelerating factor (CD55) or CD46 are not present in plaques (Zanjani, Finch et al.
2005)
although clusterin which inhibits formation of the MAC attack complex� (Fig 3) �is present in these structures (Liao, Cheng, Wang,
Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004): CD59, which inhibits formation
of the MAC attack complex (Fig 3) is also present in tangled neurones and
dystrophic neurites (McGeer, Walker et al.
1991).� In the normal brain complement receptor 1 is
specifically localised to phagocytic Kolmer cells of the choroid plexus
suggesting an important first line of defence (Singhrao, Neal,
Rushmere, Morgan, and Gasque ,1999) . However, as the herpes
simplex virus binds to this receptor (Powers, Buster, Reist,
Martin, Bridges, Sutherland, Taylor, and Scheld ,1995), it may also provide an entry
portal for cerebral viral infection.
Cytokines, immunoglobulins,
HLA-antigens, acute phase proteins and the complement pathways are all involved
in pathogen defence and the localisation of components of these systems in
amyloid plaques is consistent with a pathogenic relationship to plaque
formation. �The presence of the MAC complex in neurones might also
suggest that neuronal lysis by the MAC complex could contribute to neuronal
cell death.
The immune
system, herpes simplex and other pathogens.
HSV-1 DNA is found in many healthy brains
(Jamieson, Maitland et al. 1992)
, and the virus is obviously
able to live in harmony with the host in cerebral tissue, without provoking
cell death. Indeed the viral latency transcript may even have beneficial effects
as it inhibits apoptosis and promotes neurite sprouting in neuroblastoma cells
(Li, Carpenter et al. 2010)
. Upon reactivation, the ability of HSV-1 to inhibit the
complement pathways (see above) is likely to protect it, and the neuronal
cells in which it is harboured, from destruction. HSV-1 infection can per
se produce entorhinal cortex and hippocampal cell loss along with memory deficits , in mice
(Armien
et al,2009)
. In man, it may be necessary to� disrupt the harmony between the virus and the
host and it is possible that activation of the immune system by other pathogens
implicated in Alzheimer�s disease (e.g. Helicobacter Pylori, Chlamydia Pneumoniae
and others
(Honjo, van Reekum et al. 2009)
) might disturb this fragile
balance, allowing viral destruction at the expense of neuronal loss. Chlamydia
Pneumoniae antibodies have recently been identified in the Alzheimer�s disease
brain, colocalising in apposition to plaques and tangles in vulnerable brain
regions
(Hammond, Hallock et al. 2010)
. ��Immune activation, while promoting latent rather
than active infection can also produce collateral neuronal damage via inflammatory
mediators
(Conrady, Drevets et al. 2010)
. Paradoxically, while the
immune system is suppressed, the virus and the host cells could peacefully
coexist, while immune activation and viral destruction might be at the terrible
cost of neuronal death.�
��
APP
processing and herpes Simplex.
(Fig 5)
�
Gamma-secretase plays a key role in Alzheimer�s disease, cleaving the amyloid precursor protein APP to form the toxic beta amyloid peptide or a non-toxic intracellular signalling peptide, the APP intracellular domain (AICD). A component of this enzyme complex, Nicastrin, is upregulated by HSV-1 infection in cell culture, and beta amyloid deposition is also observed in this model, and in the brains of HSV-1 infected mice (Wozniak, Itzhaki et al. 2007). Gamma secretase also cleaves the herpes virus receptors syndecans 1 and 2 (Hemming, Elias, Gygi, and Selkoe ,2008).
The intracellular processing of full length APP follows the same route as that of the virus.� The anterograde transport of APP and HSV-1 (towards the cell surface) requires Heavy chain Kinesin-1 which binds to both APP (Szodorai, Kuan et al. 2009) and to HSV-1. Furthermore APPBP2 (Pat1), an APP binding protein, is a kinesin light chain binding protein that also binds to HSV-1 and is involved in the anterograde transport of both APP (Zheng, Eastman et al. 1998) and the virus. �Anterograde APP transport requires the viral binding protein RAB3A GTPase and the APP transport vesicle contains the viral binding synaptosomal protein SNAP25 (Szodorai, Kuan, Hunzelmann, Engel, Sakane, Sasaki, Takai, Kirsch, Muller, Beyreuther, Brady, Morfini, and Kins ,2009). APP is directly involved in the anterograde transport of the virus (in squid axons) and is a major component of the viral particles (Satpute-Krishnan, DeGiorgis, and Bearer ,2003).
APP endocytosis, like that of HSV-1 is mediated by dynamins, including the viral binding DNM2, and is regulated by the viral binding protein RAB5A (Marquez-Sterling, Lo et al. 1997). Decreasing DNM2 levels increases beta-amyloid secretion in neuroblastoma cells (Kamagata, Kudo et al. 2009). Such an effect could theoretically be duplicated by viral binding to DNM2. The retrograde transport of APP, as for HSV-1, involves dyneins and dynactins, and dynein intermediate chain knockdown increases intracellular APP and tau levels in neuroblastoma cells (Kimura, Imamura et al. 2007)
The gamma secretase mediated cleavage of APP results in the formation of� beta-amyloid and also of an intracellular signalling peptide, APP intracellular domain (AICD), which complexes with Fe65 (APPB1) and Tip60 (KAT5) or with APPB1 and TFCP2, creating active transcription complexes that increases the expression of various target genes, including APP, the APP beta-secretase BACE1 and GSK3B (Tip 60 complex) or of GSK3B (TFCP2 complex), �effects accompanied by increased� phosphorylation of the microtubule associated protein tau ��(Muller, Meyer et al. 2008) �The nuclear viral binding protein Lamin A/C binds to Tip60 (Stelzl, Worm et al. 2005)and TFCP2 bind to the viral origin of replication site (Dabrowski, Carmillo et al. 1994). AICD is degraded by caspase 3 (Chow, Mattson et al. 2009).
Following its entry into
the nucleus the HSV-1 viral protein ICP0 localises in nuclear dot 10 (ND10)
bodies. Its ubiquitin E3 ligase activity, in concert with the host ubiquitin E2
conjugating enzyme UBE2D1, degrades the PML and SP100 components of the ND10
body (Gu and Roizman ,2009). PML binds to TIP60, recruiting it to PML nuclear bodies (Wu, Hu et al. 2009). The major susceptibility gene PICALM, binds to an
exportin involved in nuclear viral egress (see below) and is thus in a position
to regulate the viral effects on PML3 and tip60, and APP signalling.
Thus, as shown in Fig 5,
the virus is clearly in a position to regulate APP processing, intracellular signalling
and tau phosphorylation which in turn
are potentially able to influence viral DNA production and the transport of the
virus along the microtubule network.
Beta-amyloid processing and herpes simplex
(Fig 5)
Fibrillar beta-amyloid, the toxic form of the peptide, enhances the infectivity of several viruses including herpes simplex in various cell lines. It was suggested that this might be due to enhanced fusion of the lipid envelope of these viruses with the cell membrane as the amyloid peptides and other artificial fibril producing peptides also promoted the association of lipid vesicles with cells (Wojtowicz, Farzan et al. 2002). In� addition amino acids 1-42 of HSV-1 glycoprotein B show 33% amino acid identity with beta-amyloid (Pyles ,2001), raising the possibility of viral-induced autoimmunity to beta-amyloid, a process that could continue after elimination of the virus.�� Indeed, beta-amyloid autoantibodies have been found in elderly patients and AD patients , although viral cross-reactivity has not been assayed (Sohn, So et al. 2009). A peptide from the homologous region of glycoprotein B, like beta-amyloid, forms fibrillar aggregates, accelerated beta-amyloid fibril formation� and displays cytotoxic properties superior to those of beta-amyloid (Pyles ,2001).
Lipoprotein receptors play a key role in the clearance of potentially toxic beta-amyloid. The principal routes are mediated by the lipoprotein receptor LRP1, which binds to A2M- or APOE-bound beta-amyloid� and by LRP2 and LRP8 which bind to clusterin-bound beta-amyloid. Beta amyloid is also degraded by the viral binding insulin degrading enzyme (IDE) (Bates, Verdile et al. 2009). The APOA1 lipoprotein transporter, ABCA1 is also involved in beta-amyloid clearance (Koldamova, Staufenbiel et al. 2005).
The HSV-1 binding protein, complement C3 is
also a ligand for LRP1 and LRP8, both of which play a role in C3 cellular
uptake (Meilinger, Gschwentner
et al. 1999).
Beta amyloid in the bloodstream is processed by its binding to C3, which
subsequently binds to complement receptor 1 on erythrocytes(Rogers, Li et al. 2006). Viral binding to A2M, C3, APOA1,
or APOE as well as to IDE suggest multiple means by which the virus might influence
beta-amyloid clearance as shown in Fig 5. The effects of clusterin, which
inhibits the formation of the MAC attack complex by binding to several of its
components are also shown. These compartments may well �be linked as Megalin (LRP2) antibodies
induce a marked activation of the MAC complex in kidney subepithelial cells,
suggesting that impaired clusterin import via LRP2� reduces its inhibitory effects on MAC complex
formation� (Ronco and Debiec ,2007).
Thus, many components the APP physiological and toxic network are also involved in the herpes simplex life-cycle and the virus binds to several key components involved in full length APP processing, APP signalling and beta-amyloid processing. These networks are also related to GSK3B expression and tau phosphorylation, and these relationships perhaps explain why HSV-1 infection leads to increased beta-amyloid deposition and tau phosphorylation. Similarly these APP processing networks are likely to impinge upon viral function.
Mitochondrial gene deletion �(Fig 1, Fig
2)
Mitochondrial gene deletion is a feature in the Alzheimer�s disease brain although this can also be observed in normal subjects (Corral-Debrinski, Horton et al. 1994). However, increased levels of mitochondrial DNA deletion, relative to controls, have been observed in the cerebral vasculature in Alzheimer�s disease (Aliyev, Chen et al. 2005) . Mitochondrial gene deletion can be produced by the herpes viral protein UL12.5, which enters mitochondria, deleting mitochondrial DNA (Saffran, Pare et al. 2007). As mentioned above, mitochondria cluster around the virus during infection and a large number of amyloid plaque related proteins are of mitochondrial origin.
�
Major susceptibility
genes and herpes simplex
APOE, clusterin, complement receptor
1 and phosphatidylinositol binding clathrin assembly protein, PICALM are the
major genetic risk factors in Alzheimer�s disease (see http://www.polygenicpathways.co.uk/alzpolys.html
for references). �Each of these is
related to herpes simplex. The virus binds to both APOE and complement receptor
1. It also binds to the mannose- 6-phosphate receptor M6PR, a clusterin
receptor whose endosomal routing is controlled by PICALM. The herpes simplex virus
also uses Crm1 (exportin 1, XPO1) �dependent pathways for nuclear egress, of
both viral RNAs and the HSV-1 protein UL47 (Williams, Verhagen et
al. 2008). PICALM and other endocytic-regulatory proteins bind to
Crm1 and it has been suggested that these might control the nuclear
localisation of transcription factors (Vecchi, Polo et al.
2001).
Thus, all of the major Alzheimer�s disease genetic risk factors can be implicated in the viral life-cycle. 28 lesser susceptibility genes are also directly related to the herpes simplex life cycle. 32 minor susceptibility genes are related to immune defence, and 35 susceptibility genes are related to cholesterol and lipoprotein function, an important factor, as herpes simplex viral entry in Vero cells is cholesterol and lipid raft dependent, blocked by the cholesterol synthesis inhibitor , nystatin, and restored by the addition of cholesterol (Bender, Whitbeck et al. 2003) (Table 2).
It is plausible that polymorphisms
in CR1, clusterin and PICALM, and many other genes related to herpes simplex,
might also influence viral virulence. As HSV-1 seropositivity has been reported
in 66% of the American population over the age of 12� (Schillinger, Xu et al. 2004) and in 90% of
the population over 70 (Smith and Robinson ,2002) , and viral DNA has also
been detected in the brains of patients without Alzheimer�s disease (Jamieson,
Maitland, Wilcock, Yates, and Itzhaki ,1992) , clearly� any ability to contribute to Alzheimer�s disease
has to be conditioned � perhaps by various Alzheimer�s disease susceptibility
genes, or other environmental or infection-related �risk factors. In this context, Helicobacter pylori
infection is a major cause of gastritis and also linked to duodenal and gastric
ulcers. Its effects on gastritis are both genetically and environmentally
conditioned. H.Pylori afflicts a substantial proportion of the world population
(~50%), not all of whom develop gastrointestinal problems (Brown
,2000) .
ADNP activity-dependent
neuroprotector homeobox .
ADNP is a neuroprotective peptide active at femtomolar concentrations against a variety of toxic insults including glutamate receptor overactivation and beta-amyloid . ADNP stabilises microtubules, and peptide derivatives of ADNP are in clinical trials and have already shown promise in mild cognitive impairment (Gozes, Stewart et al. 2009). The HSV-1 protein ICP8 binds to ADNP, and ADNP sequestration may be an important hub of the viral toxic effects. Blockade of this interaction might be considered as a therapeutic option.
HSV-1 Proteins in the
Alzheimer�s disease brain.��
Detection of HSV-1 proteins in Alzheimer�s disease has seldom
been studied .The limited data are recorded here. HSV-1 immunoreactivity was
detected in one Alzheimer�s disease subject, and also in one control in a
During its sojourn in the brain, the virus exists predominantly in a latent state, where few viral proteins are expressed. Indeed, during this state the virus may exert protective effects via inhibition of apoptosis and the promotion of neurite growth (Li, Carpenter, Hsiang, Wechsler, and Jones ,2010) . However, reactivation from time to time may occur, that could reignite the host/pathogen battle. Factors able to reactivate the virus from this dormant state include cytokines and growth factors� and 17-beta estradiol (Kriesel ,1999;Laycock, Brady et al. 1994) (Vicetti Miguel, Sheridan et al. 2010) . It has also been suggested that immune activation by the virus, while driving it to a latent state and inhibiting replication, produces collateral damage via the activation of inflammatory, and toxic, mediators and the production of toxic free radicals (Conrady, Drevets, and Carr ,2010) .�� The establishment of latency and the expression of viral immediate early genes is influenced by APOE4 (Miller and Federoff ,2008).
One could also argue that the
complement and immune systems have successfully eliminated the antigenic viral
constituents, leaving behind the non-antigenic cellular proteins with which it
was associated.
Any such successful elimination or suppression of the virus might have been at the cost of destruction of the cells containing the virus, a scenario that may be supported by the presence of numerous immune related proteins in plaques and tangles and the demonstration of the membrane attack complement complex in Alzheimer�s diseases dystrophic neurites and in the neuronal cytoplasm (McGeer, Akiyama, Itagaki, and McGeer ,1989;Itagaki, Akiyama, Saito, and McGeer ,1994).
Discussion.
There is evidence that HSV-1 infection is a risk factor in Alzheimer�s disease and infection has been reported to induce beta-amyloid deposition and tau phosphorylation in animal models (Wozniak, Itzhaki, Shipley, and Dobson ,2007;Wozniak, Frost, and Itzhaki ,2009 Refs) . Herpes simplex infection in mice also causes cortical and hippocampal neuronal loss, cerebral shrinkage and memory deficits, as observed �in Alzheimer�s disease (Armien et al,2009) . �HSV-1 DNA is also found in amyloid containing plaques in Alzheimer�s disease (Wozniak, Mee, and Itzhaki ,2009) , although evidence for viral proteins is clearly lacking. However, this may reflect the possibility that the periods of viral reactivation may be brief, and that destruction of the virus, and its proteins, has been successful. The survey of HSV-1 interacting proteins in relation to their presence in amyloid-containing plaques and tangles in Alzheimer�s disease showed a highly significant enrichment of the known HSV-1 binding proteins in these structures. 40% of 338 known viral associated proteins or chemicals are present in amyloid plaques or neurofibrillary tangles.
These plaque and tangle related proteins are general cellular constituents and the viral relationships could simply reflect the use of many cellular compartments by the virus. However, the statistical analysis suggests a highly significant enrichment of HSV-1 binding proteins in plaques and tangles, and of plaque and tangle related proteins in the HSV-1 dataset. In addition, this analysis was based on exact protein matches between the various datasets. There are, for example, multiple actins, ADP ribosylation factors, annexins, heterogeneous ribonucleoproteins, integrins, RAB�s, heat shock or 14-3-3 proteins, but those found in plaques or tangles are precisely the ones that bind to the virus. Furthermore, other factors such as mitochondrial gene deletion, as well as mitochondrial clustering around the virus and the stimulation of glucose consumption by the virus, reflected by the presence of many glycolytic enzymes and mitochondrial elements in plaques, add weight to a viral implication in the disease process. The HSV-1 binding proteins in plaques and tangles are specifically related to APP processing and tau phosphorylation respectively (Figs 4 and 5) The viral associated proteins are clearly able to affect APP processing and signalling and well as beta-amyloid processing and tau phosphorylation (Fig 3). This is indeed the case in neuroblastoma cells and in the brains of infected mice. The ordering of the human binding proteins with respect to the viral layers (envelope, virion, capsid, tegument, DNA) also suggests physiological relevance, perhaps reflecting destruction of the outer structural viral layers. �Finally, the viral associated proteins in plaques or tangles etch out a very clearly defined pathway, relating to all steps of the viral life cycle, an effect difficult to relate to chance.
The proteins present in these structures trace out the entire life cycle of the virus ranging from viral carriage in extracellular compartments, through receptors, endocytosis, intracellular vesicular traffic, mitochondrial gene deletion, nuclear transport, nuclear DNA synthesis, protein translation and quality control, the bypass of autophagy, apoptosis and protein destruction, inhibition of the complement and other defence systems, retrograde and anterograde transport and finally to exocytosis. �Most viral receptors and carriers, as well as the majority of proteins involved in HSV-1 transport, and most of the host proteins incorporated as HSV-1 virion components are also present in amyloid containing plaques. Viral associated proteins are also components of neurofibrillary tangles, although the class of proteins involved are more restricted to carriers, receptors, anterograde and retrograde transport and kinases/phosphatases involved in tau phosphorylation.
Throughout these networks there are abundant relationships between the virus and APP or tau processing, the core pathological deficits in Alzheimer�s disease. The virus and APP use the same kinesin motors and microtubule network for transport to the cell surface, beta-amyloid facilitates the entry of HSV-1 and other enveloped viruses, and hijack of the ubiquitin proteasome network by the virus is likely to interfere with beta-amyloid processing. The virus also binds to components of the intracellular APP signalling network and to many of the components that are crucial for beta-amyloid clearance.
The use of the microtubule network as a railway track for HSV-1 transport suggests a relationship with microtubule dynamics that is supported by the ability of viral infection to promote tau phosphorylation (Wozniak, Frost, and Itzhaki ,2009;Zambrano, Solis et al. 2008) . Many of the viral associated proteins present in tangles are related to tau phosphorylation.
The implication of many Alzheimer�s disease susceptibility genes in the viral life cycle, or in immunity, suggest that viral involvement might be genetically conditioned.��
The concentration of viral- and immune related proteins in plaques and tangles, and the presence of the complement membrane attack complex in neurones suggests that plaques and tangles represent cemeteries for a battle between the virus and the host�s immune defence mechanisms, which may well have been won, but at the terrible cost of �extensive complement-mediated neuronal loss.�
.����������
In summary, the high proportion of viral associated proteins in amyloid containing plaques and/or NFT�s supports accumulating evidence for the involvement of HSV-1 in the pathology of Alzheimer�s disease. Therapies directed towards the immune network or the complement system might therefore be of benefit in this condition. Blockade of the interaction between the neuroprotective peptide ADNP, and the viral protein ICP8, may also be envisaged. In addition, as the virus is essentially cytoprotective in its latent state, blocking apoptosis and promoting neurite extension in neuroblastoma cells (Li, Carpenter, Hsiang, Wechsler, and Jones ,2010), factors that drive the virus to latency, or prevent its reactivation may well be beneficial. As already suggested (Wozniak, Mee, and Itzhaki ,2009;Wozniak, Frost, and Itzhaki ,2009), vaccination against HSV-1 or antiviral therapies in the early stages of Alzheimer�s disease could be considered as viable therapeutic options in Alzheimer�s disease.
Acknowledgements: I would like to thank Oliver Chao and Nasire Mahmudi for help in obtaining reprints, the authors who provided information for the HSV-1 database and Bob Blizard for statistical advice.
Table 1: Statistical
analysis of the HSV-1 binding proteins present in plaques or tangles (A) and of
plaque and tangle components found in the HSV-1 binding protein dataset (B).
Chi squared and p values are provided.
A)Enrichment of HSV-1 binding proteins, (VBP�s), in the plaque or tangle proteomics datasets. Expected values are based on a total of 27478 contigs in the human genome http://www.ensembl.org/Homo_sapiens/Info/Index. The plaque proteome also contained tubulins (N=2), H1-H4 histones (N=4) and the chondroitin sulphate proteoglycans, appican (APP), neurocan and Versican which are potential viral associated proteins, while dynamins, dyneins and kinesin (N=4) in these structures are involved in viral transport. Tubulins (N=4), H1 or H2 histones (N=11) and the chondroitin sulphate proteoglycan, Versican, were also components of the tangle proteome and are potential viral binding partners and/or involved in viral transport. Calculations with or without these proteins are included.
|
Enrichment of HSV-
binding proteins in plaques or tangles |
Number of VBPs |
Number of proteins
in plaque/tangle� dataset |
Expected percentage
of VBP�s in any dataset (338/27478)% |
Observed percentage of VBP�s |
Expected number of
HSV-1 interacting proteins in dataset |
Observed number, of
HSV-1 proteins in� dataset |
Fold enrichment (By N) |
Chi squared DF=1 |
P value |
|
Plaques |
338 |
488 |
1.23 |
13.1 |
6 |
64 |
10.7 |
567 |
2.5229E-125 |
|
Tangles |
338 |
79 |
1.23 |
17.7 |
1 |
14 |
14.4 |
171 |
4.47466E-39 |
|
Plaques
Including potential partners |
338 |
488 |
1.23 |
15.8 |
6 |
77 |
12.8 |
851 |
4.4072E-187 |
|
Tangles
Including potential partners |
338 |
79 |
1.23 |
38.0 |
1 |
30 |
30.9 |
851 |
4.4072E-187 |
B) Enrichment of plaque or tangle components in the HSV-1 binding protein dataset. Calculations with or without potential binding partners are included.
|
Enrichment of
plaque or tangle proteins in the HSV-1 binding dataset |
Number of proteins
in plaques or tangles |
Number of VBP�s |
Expected percentage
of plaque or tangle proteins in any other dataset 488/27478% or 79/27478% |
Observed percentage |
Expected number of
plaque /tangle proteins in� HSV-1
dataset |
Observed number of
plaque/tangle proteins in HSV1 dataset |
Fold enrichment (By N) |
Chi squared DF=1 |
P value |
|
Plaques |
488 |
338 |
1.78 |
18.9 |
8.7 |
64 |
7.4 |
345 |
5.2E-77 |
|
Tangles |
79 |
338 |
0.29 |
4.1 |
0.23 |
14 |
61.6 |
169 |
9.51E-39 |
|
Plaques
Including potential partners |
488 |
338 |
1.78 |
22.8 |
8.7 |
77 |
8.9 |
527 |
1.3E-116 |
|
Tangles
Including potential partners |
79 |
338 |
0.29 |
8.9 |
0.23 |
30 |
132.1 |
843 |
2.4E-185 |
Calculation rationale: The human genome contains 27478 contigs. 338 viral associated proteins are known; 488 proteins were found in plaques and 79 in tangles in the two proteomics studies. The expected percentage of each class of protein in any other dataset is thus 338, 488 or 79 /27478 %, from which the expected number of each class of proteins in any particular dataset can be calculated. The observed and expected numbers of proteins absent from each dataset were also calculated, providing the 2*2 table for calculating Chi squared and the p values.
E.g.
|
HSV-1 binding proteins
in plaques |
Observed |
Expected |
Chi2 |
Value |
|
In
plaques |
64 |
6 |
|
|
|
Not
in plaques |
424 |
482 |
567 |
2.5229E-125 |
Table 2
HSV-1 binding
proteins (VBP) classified by function and their representation in amyloid
plaques and neurofibrillary tangles. Alzheimer�s disease susceptibility genes
directly and indirectly related to herpes simplex are also shown.�
Carriers are defined as soluble ligands occupying extracellular compartments (e.g. A2M, APOA1, APOE, and Factor XII). *HSV-1 binds to chondroitin sulphate or heparan sulphate proteoglycans, which are present in amyloid plaques, but which have not been individually tested for HSV-1 binding. Viral associated proteins are referenced in an online database at http://www.polygenicpathways.co.uk/herpeshost.html. Genes implicated in Alzheimer�s disease are referenced at http://www.polygenicpathways.co.uk/alzpolys.html
Proteins are identified by Gene symbols (See Tables 3 to 8 for names and references). Proteins in bold are specific to plaques or tangles.
|
Viral Class of
protein |
Percent in plaques |
VBP�s in plaques |
Percent in tangles |
VBP�s in tangles |
% in both |
|
Carrier (soluble ligands) |
100% (4/4) |
A2M, APOA1, APOE, FGF2 |
50% (2/4) |
APOE FGF2 |
100% (4/4) |
|
Receptors |
68% (15/22) 77% ( 23/30 with CSPG�s and HSPG�s |
Heparan sulphate Chondroitin sulphate ANK2, FGFR1, IDE, IGF2R, ITGA5, ITGB2 ITGB3, ITGAM, M6PR, MAG, SDC1, SDC2: Also LTF which blocks viral entry. |
27% (6/22) 47% (14/30) with CSPG�s and HSPG�s |
Heparan sulphate Chondroitin sulphate FGFR1, NCL, SDC1, SDC2: and LTF |
64% (21/33) 70% (28/40) with CSPG�s and HSPG�s |
|
* Chondroitin sulphate proteoglycans |
- |
APP (Appican) Neurocan, Versican |
- |
Versican |
|
|
* Heparan sulphate proteoglycans |
- |
Aggrecan, Agrin, Glypican 1, Perlecan,� Syndecans SDC3, SDC4 |
- |
Agrin, Glypican , Perlecan Syndecans SDC3, SDC4 |
|
|
Free radical |
67% (2/3) |
TXN, PRDX1 |
67% (2/3) |
PRDX1, PRDX2 |
100% |
|
Metabolic |
71% (5/7) |
ATP5J, GAPDH, NME1, TPI1, SLC25A5 |
29% (2/7) |
GAPDH SLC25A5 |
71% (5/7) |
|
Endocytosis |
82% (9/11) |
ALIX, ARF1, ARF3, ARF4, ARF5, IGF2R, M6PR, RAB10 RAB5A |
18% (2/11) |
GRB2 IGF2R |
91% (10/11) |
|
Intervesicular |
88% (7/8) |
IGF2R, M6PR, RAB2A, RAB2B, RAB33B, RAB4B RAB6A |
13% (1/8) |
IGF2R |
88% (7/8) |
|
Retrograde to nucleus |
31% (5/16) |
DCTN1, DYNC1L1,� DYNLL1, YWHAG YWHAZ |
63% (10/16) |
CDK1 CSNK2B SET tau TUBA4A TUBB2A TUBB3 TUBB3 TUBB4 TUBB4Q YWHAZ |
94% 15/16 |
|
Nuclear transport |
40% (6/15) |
ACTB, ACTG1, CFL1, CSRP1, LMNA, PFN1 |
40% (6/15) |
ACTB, ACTG1, CFL1, KPNA2, NCL, NUP358 |
60% (9/15) |
|
Actin related |
86% (6/7) |
ACTB, ACTG1, CDC42,
CFL1, CSRP1, PFN1 |
43% (3/7) |
ACTB ACTG1 CFL1 |
6/7 |
|
Anterograde to plasma membrane |
25% (4/16) |
APP, YWHAE , YWHAG,
YWHAZ |
69% (8/16) |
CDK1 CSNK2B SET tau TUBA4A TUBB2A TUBB3 TUBB3 TUBB4 TUBB4Q YWHAZ |
85% (14/16) |
|
Exocytosis |
78% (7/9) |
ANXA1, ANXA2,
ANXA5, GAP43, RAB35, RAB3A, SNAP25 |
11% (1/9) |
ANXA5 |
78% (7/9) |
|
APP processing |
72% (13/18) |
APOA1, APOE, APP, C3, CASP3, DCTN1, DYNC1L1, F12, F8, LMNA, RAB3A, RAB5A, SNAP25 |
17% (3/18) |
APOE, C3, CASP3 |
72% (13/18) |
|
Tau phosphorylation |
29% (3/7) |
PRKACA YWHAZ tau |
86% (6/7) |
CDC2, CSNK2B PRKACA SET, tau, YWHAZ |
86% (6/7 ) |
|
Coagulation factors |
50% (4/8) |
F3, F8, F12 Kallikrein |
0 |
- |
50% (4/8) |
|
Globins |
75% (3/4) |
HBA1, HBB, HBE1 |
0 |
|
75% 3 /4) |
|
Autophagy |
75% (6/8) |
BECN1 DCTN1 RAB10 RAB33B RAB35 RAB5A |
38% (3/8) |
BECN1 CHMP2B EIF2AK3 |
100% (8/8) |
|
Heat shock proteins and protein stress |
63% (5/8) |
CANX, HSPA1A ,HSPA8, HSAP1L HSP90AA1 |
50% (4/8) |
HSP40, HSPA1A ,HSPA8 EIF2AK3 (pretangles) |
86% (6/7) |
|
Mitochondrial |
75% (3 / 4) |
DNA deletion SLC25A5 Impaired respiration |
50% (2/4) |
DNA deletion SLC25A5 |
50% |
|
Immune and defence-related |
39%(14/36) |
BECN1, C3, EIF2AK2, HLA-DRB1, HLA-DRB3, HLA-DRB4, IFNGR1, �IgG , IL6, MIF, PRKRA, PPIA, PSMA2, XRCC6 |
28% (10/36) |
BECN1, CD59, EIF2AK2 EIF2AK3, HLA-DRB1, HLA-DRB3, HLA-DRB4,
IFNGR1, IL6, PPIA |
44% |
|
Apoptosis |
50% (6/12) |
BAX, CYTC1, CASP3, CST3, SMAD3 TGFB1, |
33% (4/12) |
BAX, CAPS3, CST3,� SMAD3 |
40% |
|
Translation |
18% (2/11) |
EEF1A1 EIF4A2 |
27% (3/11) |
EEF1A1 EIF2S1,
EIF4E |
30% |
|
Ubiquitin/Proteasome |
17% (2/12) |
PSMA2, UBC |
8% (1/12) |
UBC |
17% |
|
Nuclear |
14% (12/86) |
APEX1 Histone H1 Histone H2A Histone H2B Histone H4 HMGA1 HMGB1 HNRNPK JUND LMNA NFKB1 TP53 |
17% (15/86) |
BRCA1 Histone H1 Histone H2A Histone H2B Histone H4 HNRNPK JUND KPNA2 NCL NUP358 PCNA SET SP1 SP3 TP53 |
26% (/86) |
|
Replication repair |
16% (3/19) |
APEX1, PARP1, XRCC6 |
11% (2/19) |
BRCA1 MCM2 |
26% (5/19) |
|
Cell cycle |
13% (1/8) |
CDC42 |
13 % (1/8) |
CDC2 |
25% (2/8) |
|
Signalling |
33% (3/9) |
JNK, P38, PRKACA |
63% (6/9) |
CSNK2B, GRB2 SET, JNK, P38 PRKACA |
|
|
Chromatin remodelling |
0 (0/19) |
- |
5% (1/19) |
CHMP2B |
5% |
|
RNA splicing |
0 (0/5) |
- |
0 (0/5) |
|
0% |
|
Classes of viral
proteins |
|
|
|
|
|
|
Surface proteins |
79% (26/33) |
All Carriers and receptors (see above) |
48% (16/33) |
Carriers and receptors (see above) |
82% (27/33) |
|
Virion components |
63% (32/51) |
ACTB, ACTG1, APP, ARF1 ,ARF3,ARF4, ARF5 ,CFL1, CSRP1, HSPA1L, �PFN1, RAB2A, RAB2B, RAB4B, RAB5A, RAB6A, RAB7A, RAB10, RAB33B, RAB35,ANXA1,ANXA2, ANXA5, GAPDH, NME1,TPI1, PRDX1, UBC, PPIA, YWHAE, YWHAG, YWHAZ |
25% (13/51) |
ACTB, ACTG1, ANXA5, CD59, CFL1, CSNK2B GAPDH, GRB2, PPIA, PRDX1, PRDX2,� UBC, YWHAZ |
71% 36/51 |
|
Capsid binding proteins |
26% (5/19) |
ANK2 DYNLL1,EEF1A1,�
RAB3A, SNAP25 |
11% (2/19) |
EEF1A1 NCL |
25% (3/19) |
|
Tegument binding proteins |
25% (9/35) |
BAD, CASP3, DYNLL1, GAP43, IFNGR1, PRKCA, RAB3A SLC25A5, SNAP25 |
21% (8/35) |
BAD CASP3 CDK1 IFNGR1 NUP358, PRKCA SET SLC25A5 |
33% (12/35) |
|
Transcription factors and genome binding |
23% (10/43) |
Histone H1 Histone H2A Histone H2B Histone H4 HMGA1, HSPA1A, IL6, JUND, NFKB1 XRCC6 |
16% (7/43) |
Histone H1 Histone H2A Histone H2B Histone H4 JUND SP1 SP3 |
14% |
|
Proteins binding to ICP8 (single stranded DNA binding protein) |
3% (1/31) |
PARP1 |
6% (2/31) |
BRCA1 MCM2 |
|
|
Alzheimer�s disease
susceptibility genes directly related to the viral life cycle. |
|||||
|
Major genes: APOE, CR1, CLU, PICALM Minor genes:
A2M, ACAN, APOA1, APP, BIN1, CST3, DNM2, EIF2AK2, gamma secretase (APH1B,
NCSTN, PSEN1, PSEN2, PSENEN), F13A1, GAPDH, GSK3B, HLA-DRB1, HSPG2, IDE, LCK,
LMNA, MAPT, MIF, NCL, PARP1, POU2F1, PVRL2, TAP2, TFCP2, TGFB1, TP53, XRCC1, �YWHAZ |
|||||
|
Immune related
susceptibility genes: RAGE
receptor AGER; Complement
components C4A, C4B, CR1; Chemokines
CCL2, CCL3, CCR2; CD molecules CD14,
CD33, CD36, CD86; C-reactive protein
�CRP; Defensin DEFB122; �Immunoglobulin receptor FCER1G; HLA-antigens HLA-A2, HLA-DRB1, MICA; Interleukins �IL10, IL18, IL1A, IL1B, IL1RN, IL33, IL6,
IL8; Lymphocyte tyrosine kinase LCK
Cyclo-oxygenase PTGS2, Cytokines CSF1, TGFB1, TNF Toll receptor TLR4 |
|||||
|
Cholesterol and
lipoprotein related susceptibility genes |
|||||
|
Cholesterol: Transporters ABCA1, ABCA2, ABCC2, ABCG1, CETP, Cholesterol metabolism CH25H, CYP46A1, DHCR24, HMGCR, HMGCS2, SOAT1, Transcription factor SREBF1 Lipoprotein: A2M,
APOA1, APOA4, APOA5, APOC1, APOC2, APOC3, APOC4, APOD, APOE, LPA Lipoprotein receptors LDLR, LRP1,
LRP2, LRP6, LRP8, LRPAP1, OLR1, VLDLR; Lipases
LIPA, LIPC,� LPL Sortilin SORL1 |
|||||
|
Functional classification of the amyloid plaque proteome according to Liao et al, VBP�s have been added. VBP+mitochondria (113); Metabolism (72); VBPs (64); Cytoskeleton (51); membrane trafficking (50); mitochondria (49); others (45) G-protein pathways (36) kinases/phosphatases and
regulators (33); proteolysis (32); Cell adhesion (26); inflammation (24);
nuclear activities (22); chaperones (16); channels/receptors (16) oxidative
stress (10); Cell death (7) |
|||||
Table 3
Primary viral receptors and carriers found in amyloid plaques or NFT�s. *HSV-1 binds to chondroitin sulphate or heparan sulphate proteoglycans, which are present in amyloid plaques, but which, apart from SDC1 and SDC2, have not been individually tested for HSV-1 binding. Viral associated proteins in this and all other tables are referenced in the online database at http://www.polygenicpathways.co.uk/herpeshost.html
|
Chemical receptors |
Localisation
in plaques and/or tangles |
|
Heparan sulphate |
Associated
with amyloid plaques (Bruinsma, te et al. 2010)and NFT�s (Perry, Siedlak et al. 1991) |
|
Chondroitin 4,6
sulphate� |
Antibodies to Chondroitin -4 or -6 sulphate label both
plaques and NFT�s in AD (DeWitt, Silver et al.
1993) |
|
Heparan sulphate
proteoglycans |
|
|
Syndecans SDC1, SDC2 |
SDC1,2,3 and 4 are found in amyloid plaques� and NFT�s(Verbeek, Otte-Holler et al. 1999) |
|
Other
receptors |
|
|
ANK2
Ankyrin 2 |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
F3
coagulation factor III (thromboplastin, tissue factor) |
Immunoreactivity
in plaques (McComb, Miller et al. 1991) |
|
F8
Coagulation factor VIII |
Occasionally
observed in neutrophils surrounding vascular and brain amyloid plaques (Savage, Iqbal et al. 1994) |
|
F12
coagulation factor XII (Hageman factor) |
Immunoreactivity
observed in senile plaques (Yasuhara, Walker et al.
1994) |
|
FGFR1 fibroblast growth factor receptor 1 |
Increased immunoreactivity in reactive astrocytes surrounding senile plaques (Takami, Matsuo et al. 1998) and in tangles (Ferrer and Marti ,1998) |
|
IGF2R insulin-like growth factor 2 receptor |
Localised to Abeta-containing neuritic plaques and in NFT�s (Kar, Poirier et al. 2006) |
|
M6PR mannose-6-phosphate
receptor (cation dependent) |
Increased
expression in pyramidal neurones (Cataldo, Barnett et al.
1997) |
|
IDE
Insulin degrading enzyme |
Present in amyloid plaques� (Bernstein, Ansorge et al. 1999) |
|
MAG
Myelin-associated glycoprotein (Suenaga, Satoh et al.
2010) |
Found
in amyloid plaques (Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
MAC-1
ITGB2 Integrin beta 2 ITGAM |
Strongly expressed in microglia in amyloid plaques (ITGB2) (Eikelenboom, Zhan et al. 1994) |
|
Vitronectin
receptor integrin alpha 5./beta 3 (ITGA5*/ITGB3*) |
Present in the microglial cores of amyloid plaques (Akiyama, Kawamata et al. 1991) |
|
Viral transporters
(ie soluble ligands that bind to receptors |
|
|
A2M
Alpha-2-macroglobulin |
Associated with amyloid plaques� (Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
APOA1 Apolipoprotein A1 |
Found in occasional senile plaques (Harr, Uint et al. 1996) |
|
APOE Apolipoprotein E |
Found in plaques, tangles and blood vessels (Namba, Tomonaga et al. 1991) |
|
FGF2 fibroblast growth factor 2 (basic) |
Present in senile� plaques and tangles (Schindowski, Belarbi et al. 2008) |
|
LTF Lactotransferrin |
Senile plaques
and tangles (Leveugle, Spik et al. 1994) |
|
Other Heparan sulphate proteoglycans* |
|
|
Aggrecan ACAN |
Present in
plaques (Bignami, LeBlanc et al. 1994) |
|
Agrin AGN |
Plaques, tangles and blood vessels (Verbeek, Otte-Holler, van den, van den Heuvel, David, Wesseling, and de Waal ,1999)) |
|
Decorin DCN |
Localised at the edges of amyloid plaques and in NFT�s (Snow, Mar et al. 1992) |
|
Glypican 1 GPC1 |
Plaques and tangles(Verbeek, Otte-Holler, van den, van den Heuvel, David, Wesseling, and de Waal ,1999), |
|
Perlecan HSPG2 |
Plaques tangles and blood vessels (Verbeek, Otte-Holler, van den, van den Heuvel, David, Wesseling, and de Waal ,1999) |
|
SDC3 |
SDC1,2,3
and 4 are found in amyloid plaques� and
NFT�s(Verbeek, Otte-Holler, van
den, van den Heuvel, David, Wesseling, and de Waal ,1999) |
|
SDC4 |
|
|
Other Chondroitin
sulphate proteoglycans* |
|
|
APP Appican |
Beta-amyloid can be generated from appican (Shioi, Pangalos et al. 1996) |
|
VCAN Versican |
Found in amyloid plaques (Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and NFT�s (Wang, Woltjer, Cimino, Pan, Montine, Zhang, and Montine ,2005) |
|
NCAN Neurocan |
Found in amyloid plaques (Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
Not found or not reported: CD209, PILRA, PVRL1, PVRL2, TNFRSF14 |
|
Table 4 HSV-1 binding proteins involved in viral transport and their presence in amyloid plaques or tangles.
|
Viral binding
protein |
Localisation in plaques and/or tangles |
|
Actins ACTB1, ACTG1 |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and of NFT�s (Wang, Woltjer, Cimino,
Pan, Montine, Zhang, and Montine ,2005) |
|
ADP-ribosylation factors ARF1 ARF3 ARF4 ARF5 ARF6 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
Annexins: ANXA1 ANXA2 ANXA3 |
|
|
ANXA5 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004). Tangle component isolated by laser dissection(Wang, Woltjer, Cimino, Pan, Montine, Zhang, and Montine ,2005) |
|
ALIX =PDCD6IP : programmed cell death 6 interacting protein |
A component of amyloid plaques (Rajendran, Honsho et al. 2006) |
|
APP Amyloid precursor protein |
Beta-amyloid� is the major plaque component(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
CANX
Calnexin |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
CDC42
cell division cycle 42 (GTP binding protein, 25kDa) |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
CFL1
Cofilin 1 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and
tangles(Wang, Woltjer, Cimino,
Pan, Montine, Zhang, and Montine ,2005) |
|
CHMP2B
chromatin modifying protein 2B |
Present
in granulovacuolar degeneration (Pretangles) (Yamazaki, Takahashi et al.
2010) |
|
CSRP1
cysteine and glycine-rich protein 1 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
DCTN1
Dynactin 1 (p150glued) |
Observed
in areas of granulovacuolar degeneration (Ateh, Hussain et al. 2008). These may be a form of autophagosome and contain
phosphorylated tau suggesting they
may precede tangle formation (Okamoto, Hirai et al. 1991) |
|
DYNC1I1� dynein, cytoplasmic 1, intermediate chain 1 |
Component
of amyloid plaques isolated by laser dissection: Plaques also contain
DYNC1H1, DYNLL2� (Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
DYNLL1 |
Component
of amyloid plaques isolated by laser dissection: Plaques also contain
DYNC1H1, DYNLL2� (Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
GAP43 growth associated protein 43 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
PFN1
Profilin 1 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
RAB
proteins members of RAS oncogene family RAB/
RAB2A / RAB2B / RAB3A/ RAB5A /RAB6A / RAB7A / RAB10 /� RAB33B / RAB35 / |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
RAB4B |
Increased
RAB4 expression in vesicles of pyramidal neurones, pre-plaque (Cataldo, Peterhoff et al.
2000) |
|
MAPT
Microtubule associated protein tau |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) Major
component of tangles. |
|
Tubulins TUBA4A, TUBB2A, TUBB3, TUBB4 and TUBB4Q |
Components of laser-dissected tangles(Wang, Woltjer, Cimino, Pan, Montine, Zhang, and Montine ,2005) |
|
SNAP25 synaptosomal-associated protein, 25kDa |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
Nuclear transport |
|
|
LMNA Lamin A |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
NCL Nucleolin |
A CDC2-phosphorylated form is found in NFT�s (Dranovsky, Vincent et
al. 2001) |
|
Dynamins, dynactins, dyneins and kinesins: Do not specifically bind to the virus but are involved in its transport along microtubules (Lyman and Enquist ,2009) |
Dynamin 1 (plaques (Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and tangles (Wang, Woltjer, Cimino,
Pan, Montine, Zhang, and Montine ,2005) )DNM1L plaques (Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004).Dynamitin (DCTN2) plaques (Ateh, Hussain, Mustafa,
Price, Gulati, Nickols, Bird, Greensmith, Hafezparast, Fisher, Baker, and
Martin ,2008) Dynactin DCTN4, Tangles (Ateh, Hussain, Mustafa,
Price, Gulati, Nickols, Bird, Greensmith, Hafezparast, Fisher, Baker, and
Martin ,2008)Other Dyneins DYNC1H1, DYNC1I1, DYNLL2, DYNLL1 Kinesin :
KIF5B (Plaques (Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) �) |
Table 5 Host proteins associated with the HSV-1 virion and their localisation in amyloid plaques or neurofibrillary tangles.
|
Viral associated
proteins |
Localisation
in plaques and/or tangles |
|
Transport related |
|
|
Actins ACTB, ACTG1 |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and of NFT�s (Wang, Woltjer, Cimino, Pan,
Montine, Zhang, and Montine ,2005) |
|
Annexins ANXA1 ANXA2 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
ANXA5� |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and of NFT�s (Wang, Woltjer, Cimino, Pan, Montine, Zhang, and Montine ,2005) |
|
ADP Ribosylation factors ARF1 ARF3 ARF4 ARF5 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
CFL1 Cofilin 1 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and of NFT�s (Wang, Woltjer, Cimino, Pan, Montine, Zhang, and Montine ,2005) |
|
CSRP1 cysteine and glycine-rich protein 1 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
PFN1 Profilin 1 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
RAB Proteins RAB2A RAB2B RAB5A RAB6A RAB7A RAB10 RAB33B RAB35 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
Heat shock |
|
|
HSPA1L |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
Metabolic |
|
|
GAPDH Glyceraldehyde 3-phosphate dehydrogenase NME1 non-metastatic cells 1, protein (NM23A) TPI1 Triosephosphate isomerase 1 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
Immune related |
|
|
CD59 CD59 antigen |
Present in tangled neurones and dystrophic neurites (McGeer, Walker, Akiyama, Kawamata, Guan, Parker, Okada, and McGeer ,1991) |
|
MIF Macrophage inhibitory factor |
A component of� amyloid plaques (Bacher, Deuster et al. 2010) |
|
PPIA Cyclophilin A |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and of NFT�s (Wang, Woltjer, Cimino, Pan, Montine, Zhang, and Montine ,2005) |
|
14-3-3 proteins |
|
|
YWHAE 14-3-3 epsilon YWHAG 14-3-3 gamma |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
YWHAZ 14-3-3 zeta |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) amd of NFT�s (Umahara, Uchihara et al. 2004) |
|
Free radical |
|
|
PRDX1 Peroxiredoxin 1 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
Ubiquitin |
|
|
UBC Ubiquitin C |
Ubiquitin is observed in both plaques and tangles although specific isoforms were not examined (He, Delaere et al. 1993)UBC is found in neurofibrillary tangles (Wang, Woltjer, Cimino, Pan, Montine, Zhang, and Montine ,2005) |
Table 6
HSV-1 binding Histones and Heat shock proteins and their presence in amyloid plaques or neurofibrillary tangles
|
Viral binding
Protein |
Localisation in plaques and/or tangles |
|
Histone
H1 |
H1,� H2 H3� histones (HIST1H1E, HIST2H2AB, HIST3H2BB)
are� Component of laser-dissected
plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004). H1 and H2 histones (HIST1H2AA ,
HIST1H2AB, HIST1H2AC, HIST1H2AD HIST1H2AI, HIST1H2AJ, H2AFJ,
H2AFX, HIST2H2AA4, HIST2H2AC, HIST3H2A)are associated with NFT�s isolated by
laser dissection (Wang, Woltjer, Cimino, Pan,
Montine, Zhang, and Montine ,2005) |
|
Histone
H2A |
|
|
Histone
H2B |
|
|
Histone H4 |
H4
histone HIST4H4 is a Component of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
HSP40 |
High
expression in homogenates correlates with fibrillar tau levels (Sahara, Maeda et al. 2007) |
|
HSP90AA1 heat shock protein 90kDa alpha (cytosolic), class A member 1 |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
HSPA1A heat shock 70kDa protein 1A (HSP72) |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and of NFT�s (Hamos, Oblas et al. 1991) |
|
HSPA8 heat shock 70kDa protein 8 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and of NFT�s (Wang, Woltjer, Cimino, Pan, Montine, Zhang, and Montine ,2005) |
Table 7 Viral binding components related to the complement and immune systems and their presence in amyloid plaques or neurofibrillary tangles.
|
Viral binding
protein |
Localisation in plaques and/or tangles |
|
C3 Complement
C3 component |
Complements C1q, C3b, C3c, C3d and C4 are components of amyloid plaques in Alzheimer�s disease (Eikelenboom and Stam ,1982) and C3 is observed in some tangles (McGeer, Akiyama, Itagaki, and McGeer ,1989) |
|
BECN1
Beclin 1 |
Found in endothelial cells in the vicinity of amyloid plaques and in dystrophic neurites, correlating with phosphorylated tau (Ma, Huang et al. 2010). |
|
HSV-1 glycoprotein E is an Fc receptor mimic that binds to the Fc domain of Immunoglobulin G (IgG) preventing its binding to complement component C1 (Friedman ,2003) |
Immunoglobulin G is located in� the amyloid plaque corona (Eikelenboom and Stam ,1982). |
|
CD59 CD59 molecule |
Present in tangled neurones and dystrophic neurites (McGeer, Walker, Akiyama, Kawamata, Guan, Parker, Okada, and McGeer ,1991) |
|
EIF2AK2 eukaryotic translation initiation factor 2-alpha kinase 2 (PKR: double stranded RNA activated protein kinase) |
PKR immunoreactivity is associated with neuritic plaques and pyramidal neurons� Its distribution correlates with that of� phosphorylated tau (Peel and Bredesen ,2003) |
|
PRKRA protein kinase, interferon-inducible double stranded RNA dependent activator (activates EIF2AK2) |
Present in
amyloid plaques (Page, Rioux et al. 2006) |
|
HLADRB1
major histocompatibility complex, class II, DR beta 1 DRB4
beta 4 |
HLA-DR molecules are found in activated microglia around plaques and tangles (Perlmutter, Scott et al. 1992) |
|
IFNGR1 interferon gamma receptor 1 |
Increased expression in astrocytes in affected areas (Hashioka, Klegeris et al. 2009) |
|
Kallikrein KLK1 |
Immunoreactivity associated with neuritic plaques (Bernstein ,1997) |
|
MIF Macrophage inhibitory factor |
A component of� amyloid plaques (Bacher, Deuster, Aljabari, Egensperger, Neff, Jessen, Popp, Noelker, Reese, Al Abed, and Dodel ,2010) |
|
PPIA
cyclophilinA |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
PSMA2� proteasome (prosome, macropain)
subunit, alpha type, 2 |
|
|
XRCC6
X-ray repair complementing defective repair 6 (Ku antigen) |
Table 8 Diverse HSV-1 binding proteins and their presence in amyloid plaques or neurofibrillary tangles.
|
Viral associated
proteins |
Localisation
in plaques and/or tangles |
|
Apoptosis related |
|
|
BAD
BCL2-associated agonist of cell death |
Increased expression in brain homogenates (Kitamura, Shimohama et al. 1998) |
|
BAX
BCL2-associated X protein |
Strongly expressed in plaques and tangles (MacGibbon, Lawlor et al. 1997) |
|
CASP3
Caspase 3 |
Increased expression in plaques and tangles (Su, Zhao et al. 2001) |
|
CST3
Cystatin C |
Localises with beta-amyloid and occasionally in� tangles (Levy, Sastre et al. 2001) |
|
CYTC1
Cytochrome C |
CYS (cytochrome c, somatic) and CYC1 cytochrome c-1 present in amyloid plaques� (Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
TGFB1
transforming growth factor, beta 1 |
Present in amyloid plaques (van der Wal, Gomez-Pinilla et al. 1993) |
|
SMAD3
SMAD family member 3 |
Present in amyloid plaques� and NFT�s (Ueberham, Ueberham et al. 2006) |
|
Metabolic |
|
|
ATP5J ATP synthase, H+ transporting, mitochondrial F0 complex, subunit F6 |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
GAPDH Glyceraldehyde 3-phosphate dehydrogenase.� |
A component of NFT�s (Wang, Woltjer, Cimino, Pan, Montine, Zhang, and Montine ,2005) and of amyloid plaques (Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
NME1 NME2 Nucleoside diphosphate kinase A/B |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
SLC25A5 solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 5 |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004). Component of�
Neurofibrillary tangles isolated by laser dissection (Wang, Woltjer, Cimino, Pan,
Montine, Zhang, and Montine ,2005) |
|
TPI Triose phosphate isomerase |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
Free radical related |
|
|
PRDX1 Peroxiredoxin 1 |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and tangles (Wang, Woltjer, Cimino, Pan,
Montine, Zhang, and Montine ,2005) |
|
PRDX2
Peroxiredoxin 2 |
Component
of Neurofibrillary tangles isolated by laser dissection (Wang, Woltjer, Cimino, Pan,
Montine, Zhang, and Montine ,2005) |
|
TXN
Thioredoxin |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
Haemoglobins |
|
|
HBA1
Haemoglobin alpha HBB
Haeomoglobin beta HBE1
Haemoglobin epsilon |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
Translation initiation |
|
|
EEF1A1 eukaryotic translation elongation factor 1 alpha 1 |
A component of NFT�s (Wang, Woltjer,
Cimino, Pan, Montine, Zhang, and Montine ,2005) and of amyloid plaques (Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
EIF4A2 eukaryotic translation initiation factor 4A2 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
EIF4E eukaryotic translation initiation factor 4E |
High levels of phosphorylated EIF4E correlate with hyperphosphorylated tau (Li, An et al. 2004). |
|
EIF2S1 eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa |
Over-expressed in cells containing NFT�s (Ferrer ,2002) |
|
Transcription
factors and others binding to viral genome |
|
|
IL6
Interleukin 6 |
Expressed
in early stage amyloid plaques and associated with tangles (Thal, Schober et al.
1997) |
|
JUND jun D proto-oncogene (AP-1) |
Overexpressed and activated in brain homogenates (Vukic, Callaghan et al. 2009) |
|
NFKB1 nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 |
Activated in cells surrounding amyloid plaques (Ueberham, Ueberham, Gruschka, and Arendt ,2006) |
|
SP1 Sp1 transcription factor |
Expressed in NFT�s, and dystrophic neurites of senile plaques� (Santpere, Nieto et al. 2006) |
|
SP3
Sp3 transcription factor |
Highly
expressed in association with NFT�s (Boutillier, Lannes et al.
2007) |
|
STAT1
signal transducer and activator of transcription 1 |
Increased
levels in brain homogenates� (Kitamura, Shimohama et al.
1997) |
|
Nuclear proteins |
|
|
APEX1
APEX nuclease (multifunctional DNA repair enzyme) 1 |
A
component of senile plaques (Tan, Sun et al. 1998) |
|
BRCA1
breast cancer 1, early onset |
Intensely
expressed in NFT�s (Evans, Raina et al. 2007) |
|
HMGA1
high mobility group AT-hook 1 |
Highly
expressed in hippocampal pyramidal neurones in damaged areas (Manabe, Katayama et al.
2003) |
|
HMGB1
high-mobility group box 1 |
Highly
expressed in amyloid plaques (Takata, Kitamura et al.
2003) |
|
Heterogeneous ribonucloprotein K
HNRNPK |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
LMNA
Lamin A |
|
|
KPNA2
karyopherin alpha 2 (RAG cohort 1, importin alpha 1) |
Increased expression in Hirano bodies (contain actins and related cytoskeletal elements) (Lee, Ueda et al. 2006) |
|
MCM2
minichromosome maintenance complex component 2 |
A phosphorylated form is associated
with neurofibrillary tangles and dystrophic neurites (Bonda, Evans et al. 2009) |
|
NCL
Nucleolin |
A CDC2-phosphorylated form is found in NFT�s (Dranovsky, Vincent,
Gregori, Schwarzman, Colflesh, Enghild, Strittmatter, Davies, and Goldgaber
,2001) |
|
NUP358
Nucleoporin 358 |
Present
in tangle bearing neurones along with karyopherin beta 2 (Sheffield and Mirra ,2008) |
|
PARP1
poly (ADP-ribose) polymerase 1 |
Overexpressed
in healthy pyramidal neurones and also in some amyloid plaques, but not in
NFT�s (Love, Barber et al. 1999) |
|
PCNA
Proliferating cell nuclear antigen |
Highly
expressed in tangle bearing neurones (Busser, Geldmacher et al.
1998) |
|
SET
SET nuclear oncogene: phosphatase 2A inhibitor I2PP2A |
Increased
hippocampal expression related to NFT�s and tau phosphorylation (Tanimukai, Grundke-Iqbal et
al. 2005)� which is regulated by SET (Chohan, Khatoon, Iqbal, and
Iqbal ,2006) |
|
TP53
Tumor suppressor p53 |
Found
in tau positive neurites and in
dystrophic processes around amyloid plaques(de la Monte, Sohn et al.
1997) |
|
14-3-3 proteins |
|
|
YWHAE tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, epsilon |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
YWHAG �tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, gamma polypeptide |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
YWHAZ tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and of NFT�s (Umahara, Uchihara, Tsuchiya, Nakamura, Iwamoto, Ikeda, and Takasaki ,2004) |
|
Diverse |
|
|
EIF2AK3 eukaryotic translation initiation factor 2-alpha kinase 3 (PERK) |
Not
seen in plaques or tangle but increased expression in pretangle neurones with
high levels of� phosphorylated tau�
(Hoozemans, van Haastert et
al. 2009) |
|
CDC2 (=CDK1) cyclin-dependent kinase 1 |
Phosphorylates
the microtubule protein tau and is
expressed in neurones with tangle-like inclusions (Pei, Braak et al. 2002) |
|
CSNK2B Casein
kinase 2 beta |
Phosphorylates
tau (Greenwood, Scott et al.
1994)and is present in NFT�s (Baum, Masliah et al. 1992) |
|
CST3 Cystatin 3 |
Component of laser-dissected plaques(Liao, Cheng, Wang, Duong, Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) |
|
GRB2 growth factor receptor-bound protein 2 |
Expression
correlates with cytoskeletal abnormalities (tangles) (McShea, Zelasko et al.
1999)Binds to Dynamin (Vidal, Goudreau et al.
1999). |
|
JNK kinase |
Activated
in granular bodies , tangles and occasional plaques (Lagalwar,
Guillozet-Bongaarts et al. 2006) |
|
P38 MAP Kinase |
Activated
in neuritic plaques and tangles (Hensley, Floyd et al. 1999) |
|
PRKACA protein kinase, cAMP-dependent, catalytic, alpha |
Component
of laser-dissected plaques(Liao, Cheng, Wang, Duong,
Losik, Gearing, Rees, Lah, Levey, and Peng ,2004) and associated with tangles : Phosphorylates tau (Jicha, Weaver et al. 1999) |
|
UBC Ubiquitin C |
Ubiquitin
is observed in both plaques and tangles although specific isoforms were not
examined (Perry, Friedman et al.
1987)UBC was identified in a
proteomics study of NFT�s (Wang, Woltjer, Cimino, Pan,
Montine, Zhang, and Montine ,2005;He, Delaere, Duyckaerts, Wasowicz, Piette,
and Hauw ,1993) |
Figure legends
Figure 1
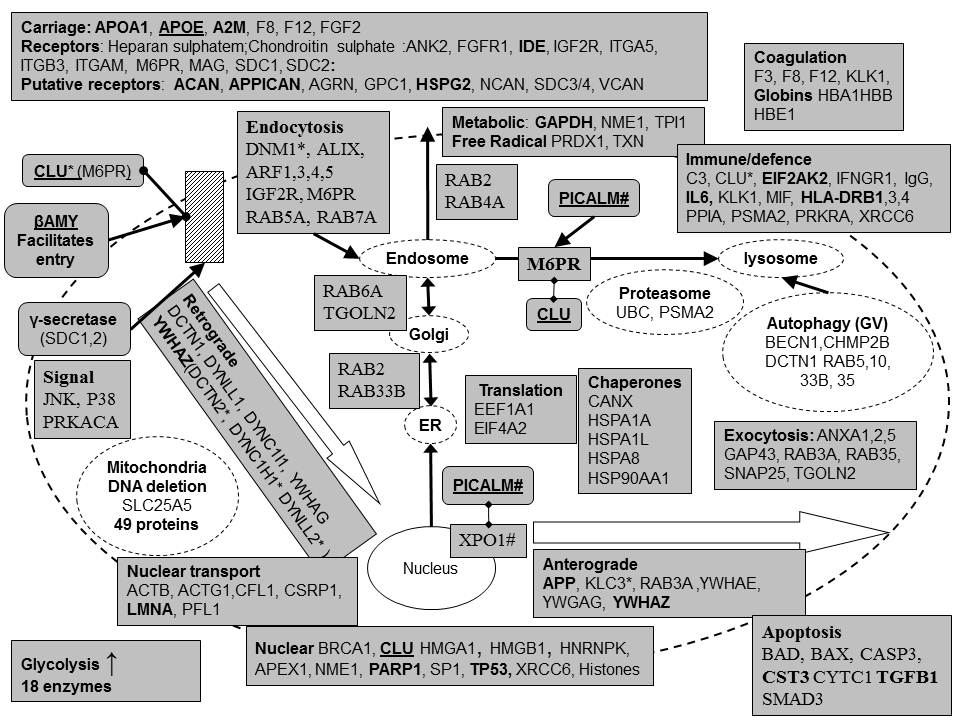
Herpes simplex binding proteins found in amyloid plaques in Alzheimer�s disease. Proteins are represented by HUGO approved gene symbols and corresponding definitions can be found in Table 3-8. All proteins in this figure bind to HSV-1 except for those marked with* (clusterin, dynamin DNM1,dynactin DCTN2, Dyneins DYNC1H1,� DYNLL2,� Kinesin KLC3, and PICALM) although these are likely to be, or are, implicated in the viral life cycle. All proteins except for those marked by # are plaque components (PICALM). Increased BAD immunoreactivity was observed in brain homogenates. 49 mitochondrial proteins and 18 glycolytic enzymes may reflect mitochondrial clustering around the virus and increased glucose consumption in infected areas (see text). Proteins are organised with respect to their cellular functions. ER = endoplasmic reticulum. βAmy = beta-amyloid. Linked diamonds (CLU/M6PR) represent binding between components. Genes in bold have been implicated as risk factors in Alzheimer�s disease and those underlined (APOE, CLU, PICALM) as major genetic risk factors (see text for details).
Figure 2
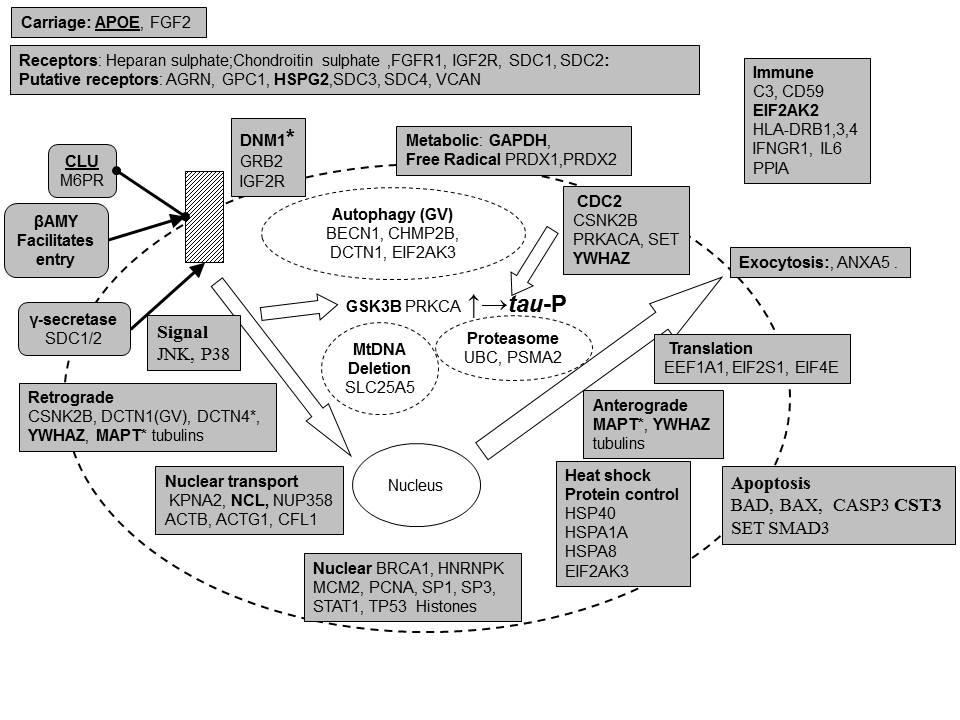
Herpes simplex binding proteins found in neurofibrillary tangles in Alzheimer�s disease. Proteins are represented by HUGO approved gene symbols and corresponding definitions can be found in Table 3-8. All proteins in this figure bind to HSV-1 except for those marked with* (DNM1, DCTN4,� MAPT (=tau) although these are implicated in the viral life cycle. All proteins except for those marked by # are recorded tangle components (PRKCA). Increased BAD immunoreactivity was observed in brain homogenates Proteins are organised with respect to their cellular functions. Genes in bold have been implicated as risk factors in Alzheimer�s disease and those underlined (APOE, CLU) as major genetic risk factors.(see text for details).
Figure 3
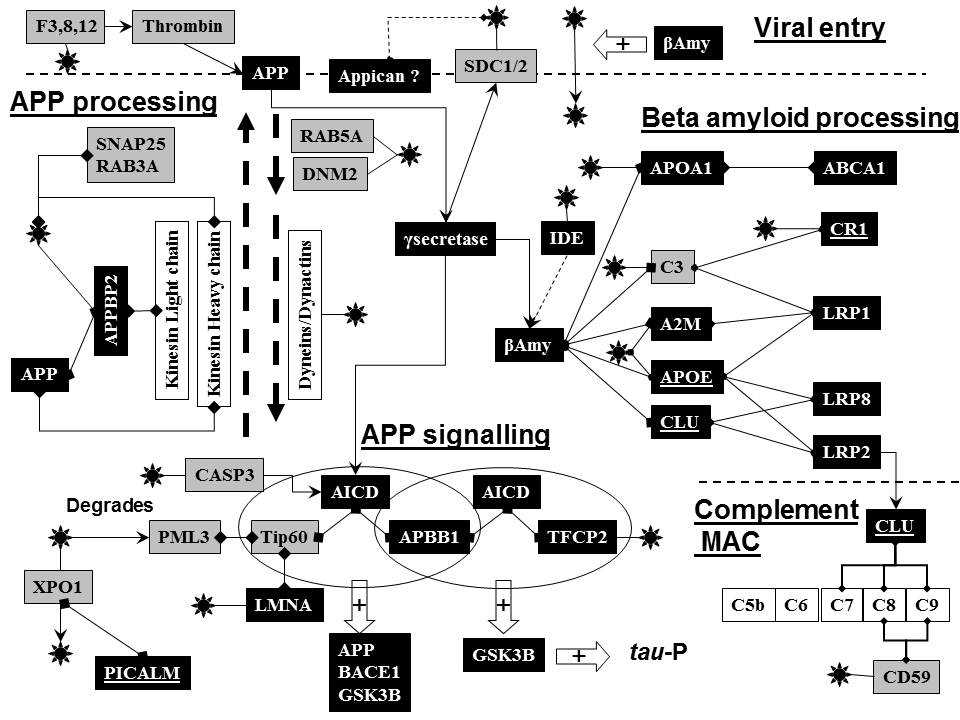
The distribution of viral associated proteins found in amyloid plaques or neurofibrillary tangles, grouped in relation to function. The classes are ranked in relation to plaque containing proteins and the number of proteins in each class is indicated on the X-axis. The types of viral component (glycoprotein, virion, capsid, tegument, Latency transcript (LAT), viral genome or ICP8) interacting with host proteins are also shown. The classification is based on Table 2, Figs 1, 2 and 5 and is detailed in the text and at to� http://www.polygenicpathways.co.uk/herpeshost.html
�.
Figure 4
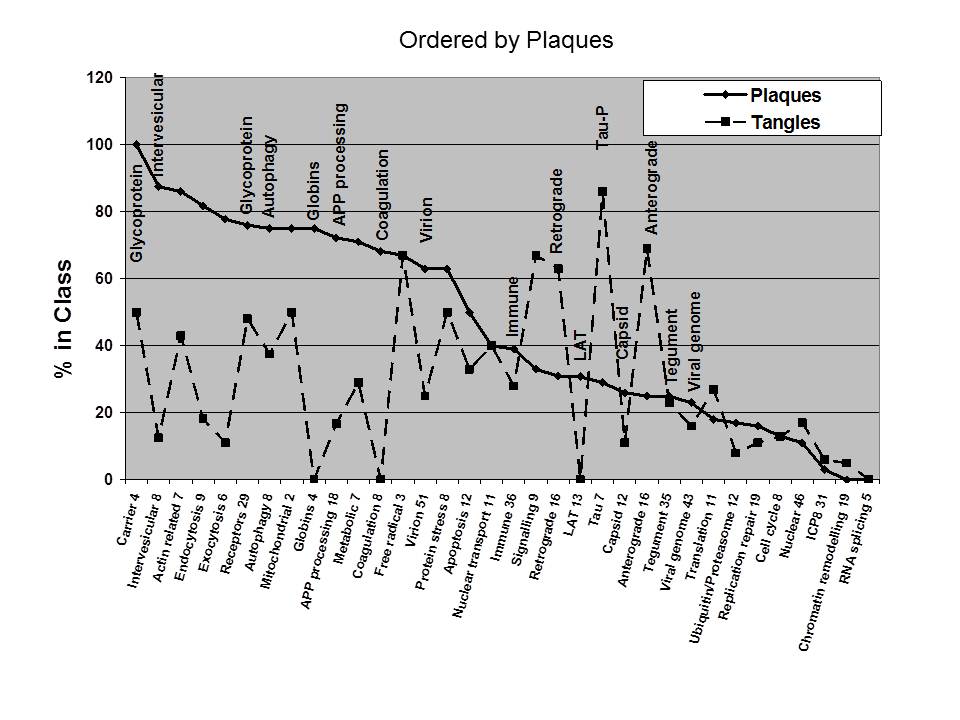
The distribution of viral associated proteins found in amyloid plaques or neurofibrillary tangles, grouped in relation to function. The classes are ranked in relation to tangle containing proteins and the number of proteins in each class is indicated on the X-axis. The types of viral component (glycoprotein, virion, capsid, tegument, Latency transcript (LAT), viral genome or ICP8) interacting with host proteins are also shown. The classification is based on Table 2, Figs 1, 2 and 5 and is detailed in the text and at to� http://www.polygenicpathways.co.uk/herpeshost.html
Figure 5
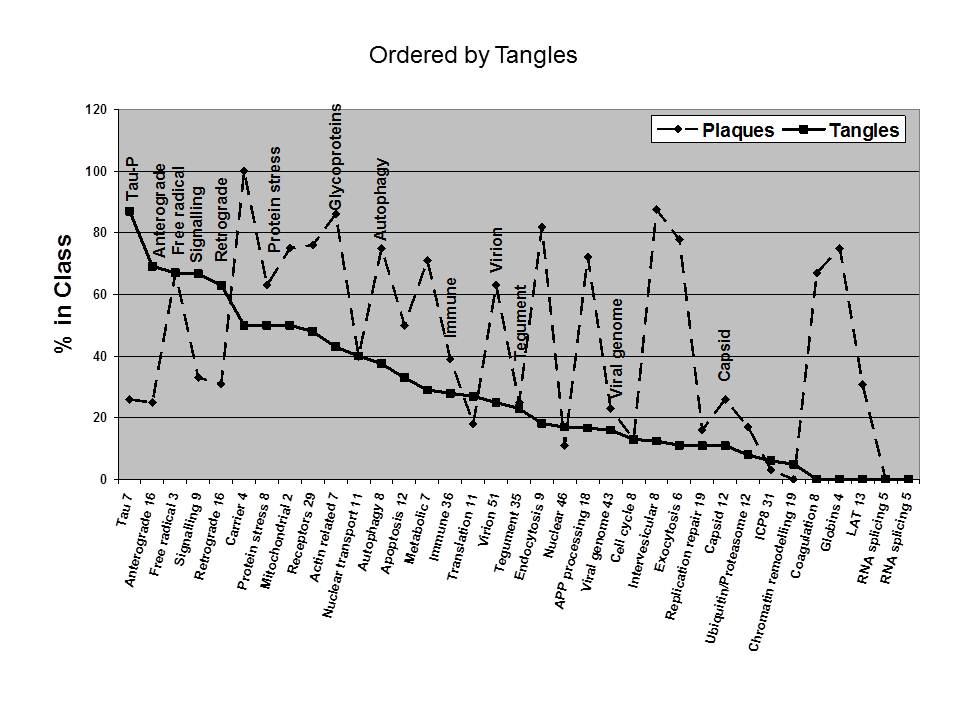
The relationships between herpes simplex and various aspects of APP physiology.
HSV-1 entry is modified by beta-amyloid (βAmy) and the APP intracellular protease complex, gamma-secretase, cleaves the viral entry receptors syndecans SDC1 and 2 (top). HSPG and CSPG (heparan and chondroitin sulphate proteoglycans, the latter including appican, an isoform of APP, are putative viral receptors. Thrombin, the coagulation factor activated protease cleaves APP.
Intracellular APP processing (de novo synthesis and transport to the plasma membrane) employs the same pathways as those used by herpes simplex and is dependent upon the viral associated proteins, SNAP25 and RAB3A. APP endocytosis is mediated via dynamin and RAB5A (top left).
Beta-amyloid (βAmy) degradation and clearance (right) is mediated by insulin degrading enzyme (IDE) and by APOE, APOA1, clusterin, and complement C3 and their respective receptors/transporters, as illustrated. Clusterin delivery via its receptor, LRP2, influences its utilisation as a complement membrane attack complex (MAC) inhibitor (bottom right).
APP intracellular signalling
(bottom) is mediated by a gamma-secretase cleaved intracellular domain of APP
(AICD) that complexes with tip60 and APBB1 or TFCP2 and APBB1, which control
the expression of APP, BACE1 and GSK3B, leading to tau phosphorylation (tau-P)
as shown. AICD is cleaved by caspase 3. HSV-1 nuclear egress is via crm1 (XPO1)
dependent pathways and PICALM binds to crm1. HSV-1 infection degrades PML3
which binds to tip60, which is also connected to the viral binding protein
lamin A (bottom left). Linked diamonds represent
binding between components and components of the HSV-1 virus that bind to the
various proteins are represented by![]() . Genes
implicated as Alzheimer�s disease risk factors are in black boxes with white
lettering and major genes (APOE, CLU, CR1 and PICALM) are underlined.
. Genes
implicated as Alzheimer�s disease risk factors are in black boxes with white
lettering and major genes (APOE, CLU, CR1 and PICALM) are underlined.
�
Akiyama, H., Kawamata, T., Dedhar,
S., and McGeer, P.L., 1991. Immunohistochemical localization of
vitronectin, its receptor and beta-3 integrin in Alzheimer brain tissue.
J.Neuroimmunol. 32, 19-28.
Aliyev, A., Chen, S.G., Seyidova,
D., Smith, M.A., Perry, G., de la, T.J., and Aliev, G., 2005.
Mitochondria DNA deletions in atherosclerotic hypoperfused
brain microvessels as a primary target for the development of Alzheimer's
disease. J.Neurol.Sci. 229-230, 285-292.
Armien, A.G., Hu, S., Little, M.R., Robinson, N.,
Lokensgard, J.R., Low, W.C., and Cheeran, M.C., 2009. Chronic
Cortical and Subcortical Pathology with Associated Neurological Deficits
Ensuing Experimental Herpes Encephalitis. Brain
Pathol.
Ateh, D.D., Hussain, I.K., Mustafa,
A.H., Price, K.M., Gulati, R., Nickols, C.D., Bird, M.M., Greensmith, L.,
Hafezparast, M., Fisher, E.M., Baker, C.S., and Martin, J.E., 2008.
Dynein-dynactin complex subunits are differentially localized in brain and
spinal cord, with selective involvement in pathological features of
neurodegenerative disease. Neuropathol.Appl.Neurobiol. 34, 88-94.
Bacher, M., Deuster, O., Aljabari,
B., Egensperger, R., Neff, F., Jessen, F., Popp, J., Noelker, C., Reese, J.P.,
Al Abed, Y., and Dodel, R., 2010. The
role of macrophage migration inhibitory factor in Alzheimer's disease.
Mol.Med. 16, 116-121.
Bates, K.A., Verdile, G., Li, Q.X.,
Ames, D., Hudson, P., Masters, C.L., and Martins, R.N., 2009.
Clearance mechanisms of Alzheimer's amyloid-beta peptide: implications for
therapeutic design and diagnostic tests. Mol.Psychiatry. 14, 469-486.
Baum, L., Masliah, E., Iimoto, D.S.,
Hansen, L.A., Halliday, W.C., and Saitoh, T., 1992.
Casein kinase II is associated with neurofibrillary tangles but is not an
intrinsic component of paired helical filaments. Brain Res. 573, 126-132.
Becker, Y., 1995. HSV-1 brain infection by the olfactory
nerve route and virus latency and reactivation may cause learning and
behavioral deficiencies and violence in children and adults: a point of view. Virus Genes. 10, 217-226.
Bello-Morales, R., Fedetz, M.,
Alcina, A., Tabares, E., and Lopez-Guerrero, J.A., 2005.
High susceptibility of a human oligodendroglial cell line to
herpes simplex type 1 infection. J.Neurovirol. 11, 190-198.
Bender, F.C., Whitbeck, J.C., Ponce,
d.L., Lou, H., Eisenberg, R.J., and Cohen, G.H., 2003.
Specific association of glycoprotein B with lipid rafts during herpes simplex
virus entry. J.Virol. 77, 9542-9552.
Bernstein, H.G., 1997. Tissue kallikrein-like
immunoreactive material decorates neuritic plaques of Alzheimer patients.
Clin.Neuropathol. 16, 69-71.
Bernstein, H.G., Ansorge, S.,
Riederer, P., Reiser, M., Frolich, L., and Bogerts, B., 1999.
Insulin-degrading enzyme in the Alzheimer's disease brain: prominent
localization in neurons and senile plaques. Neurosci.Lett. 263, 161-164.
Bignami, A., LeBlanc, A., and Perides, G., 1994. A role for extracellular matrix degradation and matrix
metalloproteinases in senile dementia? Acta
Neuropathol. 87, 308-312.
Bonda, D.J., Evans, T.A., Santocanale, C., Llosa,
J.C., Vina, J., Bajic, V.P., Castellani, R.J., Siedlak, S.L., Perry, G., Smith,
M.A., and Lee, H.G., 2009. Evidence for the progression
through S-phase in the ectopic cell cycle re-entry of neurons in Alzheimer
disease. Aging (Albany.NY). 1, 382-388.
Boutillier, S., Lannes, B., Buee,
L., Delacourte, A., Rouaux, C., Mohr, M., Bellocq, J.P., Sellal, F., Larmet,
Y., Boutillier, A.L., and Loeffler, J.P., 2007.
Sp3 and sp4 transcription factor levels are increased in brains of patients
with Alzheimer's disease. Neurodegener.Dis. 4, 413-423.
Brown, L.M., 2000. Helicobacter pylori: epidemiology
and routes of transmission. Epidemiol.Rev. 22, 283-297.
Bruinsma, I.B., te, R.L., Gevers, T., ten Dam, G.B.,
van Kuppevelt, T.H., David, G., Kusters, B., de Waal, R.M., and Verbeek, M.M.,
2010. Sulfation of heparan sulfate associated with amyloid-beta plaques in
patients with Alzheimer's disease. Acta Neuropathol.
119, 211-220.
Brunetti, C.R., Dingwell, K.S.,
Wale, C., Graham, F.L., and Johnson, D.C., 1998.
Herpes simplex virus gD and virions accumulate in
endosomes by mannose 6-phosphate-dependent and -independent mechanisms. J.Virol. 72, 3330-3339.
Bugyi, B. and Carlier, M.F., 2010. Control
of actin filament treadmilling in cell motility. Annu.Rev.Biophys. 39,
449-470.
Burch, A.D. and Weller, S.K., 2004. Nuclear
sequestration of cellular chaperone and proteasomal machinery during herpes
simplex virus type 1 infection. J.Virol. 78,
7175-7185.
Burgos, J.S., Ramirez, C.,
Guzman-Sanchez, F., Alfaro, J.M., Sastre, I., and Valdivieso, F., 2006.
Hematogenous vertical transmission of herpes simplex virus
type 1 in mice. J.Virol. 80, 2823-2831.
Burgos, J.S., Ramirez, C., Sastre,
I., Alfaro, J.M., and Valdivieso, F., 2005.
Herpes simplex virus type 1 infection via the bloodstream with apolipoprotein E
dependence in the gonads is influenced by gender. J.Virol. 79, 1605-1612.
Burgos, J.S., Ramirez, C., Sastre,
I., Bullido, M.J., and Valdivieso, F., 2002.
Involvement of apolipoprotein E in the hematogenous route of
herpes simplex virus type 1 to the central nervous system. J.Virol. 76,
12394-12398.
-----, 2003. ApoE4 is more
efficient than E3 in brain access by herpes simplex virus type 1. Neuroreport. 14, 1825-1827.
Busser, J., Geldmacher, D.S., and
Herrup, K., 1998. Ectopic cell cycle proteins predict the sites of
neuronal cell death in Alzheimer's disease brain. J.Neurosci.
18, 2801-2807.
Cardoso, C.M., Jordao, L., and Vieira, O.V., 2010.
Rab10 regulates phagosome maturation and its overexpression rescues
Mycobacterium-containing phagosomes maturation. Traffic.
11, 221-235.
Cataldo, A.M., Barnett, J.L.,
Pieroni, C., and Nixon, R.A., 1997. Increased
neuronal endocytosis and protease delivery to early endosomes in sporadic
Alzheimer's disease: neuropathologic evidence for a mechanism of increased
beta-amyloidogenesis. J.Neurosci. 17, 6142-6151.
Cataldo, A.M., Peterhoff, C.M.,
Troncoso, J.C., Gomez-Isla, T., Hyman, B.T., and Nixon, R.A., 2000.
Endocytic pathway abnormalities precede amyloid beta deposition in sporadic
Alzheimer's disease and Down syndrome: differential effects of APOE genotype
and presenilin mutations. Am.J.Pathol. 157, 277-286.
Cheshenko, N., Liu, W., Satlin, L.M., and Herold,
B.C., 2007. Multiple receptor interactions trigger release of membrane and
intracellular calcium stores critical for herpes simplex virus entry. Mol.Biol.Cell 18, 3119-3130.
Chohan, M.O., Khatoon, S., Iqbal, I.G., and Iqbal, K.,
2006. Involvement of I2PP2A in the abnormal
hyperphosphorylation of tau and its reversal by Memantine. FEBS Lett. 580, 3973-3979.
Chow, V.W., Mattson, M.P., Wong,
P.C., and Gleichmann, M., 2009. An
Overview of APP Processing Enzymes and Products. Neuromolecular.Med.
Conrady, C.D., Drevets, D.A., and Carr, D.J., 2010.
Herpes simplex type I (HSV-1) infection of the nervous system: Is an immune
response a good thing? J.Neuroimmunol. 220, 1-9.
Corral-Debrinski, M., Horton, T.,
Lott, M.T., Shoffner, J.M., McKee, A.C., Beal, M.F., Graham, B.H., and Wallace,
D.C., 1994. Marked changes in mitochondrial DNA
deletion levels in Alzheimer brains. Genomics 23, 471-476.
D'Souza-Schorey, C. and Chavrier,
P., 2006. ARF proteins: roles in membrane traffic and beyond.
Nat.Rev.Mol.Cell Biol. 7, 347-358.
Dabrowski, C.E., Carmillo, P.J., and Schaffer, P.A.,
1994. Cellular protein interactions with herpes simplex virus type 1 oriS. Mol.Cell Biol. 14, 2545-2555.
Damasio, A.R. and Van Hoesen, G.W., 1985. The limbic system and the localisation of herpes simplex
encephalitis. J.Neurol.Neurosurg.Psychiatry. 48, 297-301.
de
la Monte, S.M., Sohn, Y.K., and Wands, J.R., 1997. Correlates
of p53- and Fas (CD95)-mediated apoptosis in Alzheimer's disease.
J.Neurol.Sci. 152, 73-83.
DeWitt, D.A., Silver, J., Canning, D.R., and Perry,
G., 1993. Chondroitin sulfate proteoglycans are associated with the lesions of
Alzheimer's disease. Exp.Neurol. 121, 149-152.
Dranovsky, A., Vincent, I., Gregori, L., Schwarzman,
A., Colflesh, D., Enghild, J., Strittmatter, W., Davies, P., and Goldgaber, D.,
2001. Cdc2 phosphorylation of nucleolin demarcates mitotic stages and
Alzheimer's disease pathology. Neurobiol.Aging. 22, 517-528.
Eikelenboom, P. and Stam, F.C., 1982. Immunoglobulins
and complement factors in senile plaques. An immunoperoxidase
study. Acta Neuropathol. 57, 239-242.
Eikelenboom, P., van Exel, E., Hoozemans, J.J.,
Veerhuis, R., Rozemuller, A.J., and van Gool, W.A., 2010. Neuroinflammation
- an early event in both the history and pathogenesis of Alzheimer's disease.
Neurodegener.Dis. 7, 38-41.
Eikelenboom, P., Zhan, S.S., Kamphorst, W., van, d.,
V, and Rozemuller, J.M., 1994. Cellular and substrate
adhesion molecules (integrins) and their ligands in cerebral amyloid plaques in
Alzheimer's disease. Virchows Arch. 424, 421-427.
Evans, T.A., Raina, A.K.,
Delacourte, A., Aprelikova, O., Lee, H.G., Zhu, X., Perry, G., and Smith, M.A.,
2007. BRCA1 may modulate neuronal cell cycle re-entry in
Alzheimer disease. Int.J.Med.Sci. 4, 140-145.
Fatimathas, L. and Moss, S.E., 2010. Annexins as disease modifiers. Histol.Histopathol. 25,
527-532.
Ferrer, I., 2002. Differential
expression of phosphorylated translation initiation factor 2 alpha in
Alzheimer's disease and Creutzfeldt-Jakob's disease.
Neuropathol.Appl.Neurobiol. 28, 441-451.
Ferrer, I. and Marti, E., 1998. Distribution
of fibroblast growth factor receptor-1 (FGFR-1) and FGFR-3 in the hippocampus
of patients with Alzheimer's disease. Neurosci.Lett. 240, 139-142.
Friedman, H.M., 2003. Immune evasion
by herpes simplex virus type 1, strategies for virus survival.
Trans.Am.Clin.Climatol.Assoc. 114, 103-112.
Gozes, I., Stewart, A., Morimoto, B., Fox, A.,
Sutherland, K., and Schmeche, D., 2009. Addressing Alzheimer's disease tangles:
from NAP to AL-108. Curr.Alzheimer Res. 6, 455-460.
Greenwood, J.A., Scott, C.W., Spreen, R.C., Caputo,
C.B., and Johnson, G.V., 1994. Casein kinase II preferentially phosphorylates
human tau isoforms containing an amino-terminal insert. Identification
of threonine 39 as the primary phosphate acceptor. J.Biol
Chem. 269, 4373-4380.
Gu,
H. and Roizman, B., 2009. The two functions of herpes simplex virus 1 ICP0,
inhibition of silencing by the CoREST/REST/HDAC complex and degradation of PML,
are executed in tandem. J.Virol. 83, 181-187.
Gu,
Y., Jin, S., Gao, Y., Weaver, D.T., and Alt, F.W., 1997. Ku70-deficient
embryonic stem cells have increased ionizing radiosensitivity, defective DNA
end-binding activity, and inability to support V(D)J
recombination. Proc.Natl.Acad.Sci.U.S.A. 94, 8076-8081.
Hammond, C.J., Hallock, L.R.,
Howanski, R.J., Appelt, D.M., Little, C.S., and Balin, B.J., 2010.
Immunohistological detection of Chlamydia pneumoniae in the
Alzheimer's disease brain. BMC.Neurosci. 11, 121.
Hamos, J.E., Oblas, B.,
Pulaski-Salo, D., Welch, W.J., Bole, D.G., and Drachman, D.A., 1991.
Expression of heat shock proteins in Alzheimer's disease.
Neurology. 41, 345-350.
Harper, B.D., Beckerle, M.C., and
Pomies, P., 2000. Fine mapping of the alpha-actinin
binding site within cysteine-rich protein. Biochem.J. 350 Pt 1, 269-274.
Harr, S.D., Uint, L., Hollister, R.,
Hyman, B.T., and Mendez, A.J., 1996.
Brain expression of apolipoproteins E, J, and A-I in
Alzheimer's disease. J.Neurochem. 66, 2429-2435.
Hashiguchi, M., Sobue, K., and
Paudel, H.K., 2000. 14-3-3zeta is an effector of tau protein
phosphorylation. J.Biol Chem. 275, 25247-25254.
Hashioka, S., Klegeris, A., Schwab,
C., and McGeer, P.L., 2009. Interferon-gamma-dependent
cytotoxic activation of human astrocytes and astrocytoma cells.
Neurobiol.Aging. 30, 1924-1935.
He, Y., Delaere, P., Duyckaerts, C.,
Wasowicz, M., Piette, F., and Hauw, J.J., 1993.
Two distinct ubiquitin immunoreactive senile plaques in Alzheimer's disease:
relationship with the intellectual status in 29 cases. Acta
Neuropathol. 86, 109-116.
Hemming, M.L., Elias, J.E., Gygi,
S.P., and Selkoe, D.J., 2008. Proteomic
profiling of gamma-secretase substrates and mapping of substrate requirements.
PLoS.Biol. 6, e257.
Hensley, K., Floyd, R.A., Zheng, N.Y., Nael, R.,
Robinson, K.A., Nguyen, X., Pye, Q.N., Stewart, C.A., Geddes, J., Markesbery,
W.R., Patel, E., Johnson, G.V., and Bing, G., 1999. p38
kinase is activated in the Alzheimer's disease brain. J.Neurochem. 72,
2053-2058.
Honjo, K., van Reekum, R., and Verhoeff, N.P., 2009.
Alzheimer's disease and infection: do infectious agents contribute to
progression of Alzheimer's disease? Alzheimers.Dement. 5, 348-360.
Hoozemans, J.J., van Haastert, E.S., Nijholt, D.A.,
Rozemuller, A.J., Eikelenboom, P., and Scheper, W., 2009. The unfolded protein
response is activated in pretangle neurons in Alzheimer's disease hippocampus.
Am.J.Pathol. 174, 1241-1251.
Huemer, H.P., Menzel, H.J., Potratz, D., Brake, B.,
Falke, D., Utermann, G., and Dierich, M.P., 1988. Herpes simplex virus binds to
human serum lipoprotein. Intervirology 29, 68-76.
Igarashi, K., Murai, H., and Asaka, J., 1992. Proteolytic processing of amyloid beta protein precursor (APP) by
thrombin. Biochem.Biophys.Res.Commun. 185, 1000-1004.
Ishino, H. and Otsuki, S., 1975. Distribution
of Alzheimer's neurofibrillary tangles in the basal ganglia and brain stem of
progressive supranuclear palsy and Alzheimer's disease. Folia Psychiatr.Neurol.Jpn. 29, 179-187.
Itagaki, S., Akiyama, H., Saito, H.,
and McGeer, P.L., 1994. Ultrastructural localization of complement membrane
attack complex (MAC)-like immunoreactivity in brains of patients with
Alzheimer's disease. Brain Res. 645, 78-84.
Itzhaki, R.F., Cosby, S.L., and Wozniak, M.A., 2008.
Herpes simplex virus type 1 and Alzheimer's disease: the autophagy connection.
J.Neurovirol. 14, 1-4.
Itzhaki, R.F., Lin, W.R., Shang, D., Wilcock, G.K.,
Faragher, B., and Jamieson, G.A., 1997. Herpes simplex virus
type 1 in brain and risk of Alzheimer's disease. Lancet 349, 241-244.
Jamieson, G.A., Maitland, N.J.,
Wilcock, G.K., Yates, C.M., and Itzhaki, R.F., 1992.
Herpes simplex virus type 1 DNA is present in specific regions of brain from
aged people with and without senile dementia of the Alzheimer type. J.Pathol. 167,
365-368.
Jicha, G.A., Weaver, C., Lane, E., Vianna, C., Kress,
Y., Rockwood, J., and Davies, P., 1999. cAMP-dependent
protein kinase phosphorylations on tau in Alzheimer's disease. J.Neurosci. 19, 7486-7494.
Jin, J., Smith, F.D., Stark, C., Wells, C.D., Fawcett,
J.P., Kulkarni, S., Metalnikov, P., O'Donnell, P., Taylor, P., Taylor, L.,
Zougman, A., Woodgett, J.R., Langeberg, L.K., Scott, J.D., and Pawson, T.,
2004. Proteomic, functional, and domain-based analysis of in
vivo 14-3-3 binding proteins involved in cytoskeletal regulation and cellular
organization. Curr.Biol. 14, 1436-1450.
Kamagata, E., Kudo, T., Kimura, R.,
Tanimukai, H., Morihara, T., Sadik, M.G., Kamino, K., and Takeda, M., 2009.
Decrease of dynamin 2 levels in late-onset Alzheimer's disease alters Abeta
metabolism. Biochem.Biophys.Res.Commun. 379, 691-695.
Kar, S., Poirier, J., Guevara, J.,
Dea, D., Hawkes, C., Robitaille, Y., and Quirion, R., 2006.
Cellular distribution of insulin-like growth
factor-II/mannose-6-phosphate receptor in normal human brain and its alteration
in Alzheimer's disease pathology. Neurobiol.Aging 27, 199-210.
Kim, H.J., Lee, S., and Jung, J.U.,
2010. When autophagy meets viruses: a
double-edged sword with functions in defense and offense. Semin.Immunopathol.
Kimura, N., Imamura, O., Ono, F.,
and Terao, K., 2007. Aging attenuates dynactin-dynein interaction:
down-regulation of dynein causes accumulation of endogenous tau and amyloid
precursor protein in human neuroblastoma cells. J.Neurosci.Res.
85, 2909-2916.
Kitamura, Y., Shimohama, S., Kamoshima, W., Ota, T.,
Matsuoka, Y., Nomura, Y., Smith, M.A., Perry, G., Whitehouse, P.J., and
Taniguchi, T., 1998. Alteration of proteins regulating
apoptosis, Bcl-2, Bcl-x, Bax, Bak, Bad, ICH-1 and CPP32, in Alzheimer's
disease. Brain Res. 780, 260-269.
Kitamura, Y., Shimohama, S., Ota, T., Matsuoka, Y.,
Nomura, Y., and Taniguchi, T., 1997. Alteration of transcription factors
NF-kappaB and STAT1 in Alzheimer's disease brains. Neurosci.Lett. 237, 17-20.
Koldamova, R., Staufenbiel, M., and
Lefterov, I., 2005. Lack of ABCA1 considerably decreases brain ApoE level
and increases amyloid deposition in APP23 mice. J.Biol Chem. 280, 43224-43235.
Kosik, K.S., 1990. Tau protein and
neurodegeneration. Mol.Neurobiol. 4, 171-179.
Kriesel, J.D., 1999. Reactivation of herpes simplex
virus: the role of cytokines and intracellular factors. Curr.Opin.Infect.Dis.
12, 235-238.
Lagalwar, S., Guillozet-Bongaarts,
A.L., Berry, R.W., and Binder, L.I., 2006.
Formation of phospho-SAPK/JNK granules in the hippocampus is an early event in
Alzheimer disease. J.Neuropathol.Exp.Neurol. 65, 455-464.
Lambert, J.C., Heath, S., Even, G., Campion, D.,
Sleegers, K., Hiltunen, M., Combarros, O., Zelenika, D., Bullido, M.J.,
Tavernier, B., Letenneur, L., Bettens, K., Berr, C., Pasquier, F., Fievet, N.,
Barberger-Gateau, P., Engelborghs, S., De Deyn, P., Mateo, I., Franck, A.,
Helisalmi, S., Porcellini, E., Hanon, O., de Pancorbo, M.M., Lendon, C.,
Dufouil, C., Jaillard, C., Leveillard, T., Alvarez, V., Bosco, P., Mancuso, M.,
Panza, F., Nacmias, B., Bossu, P., Piccardi, P., Annoni, G., Seripa, D.,
Galimberti, D., Hannequin, D., Licastro, F., Soininen, H., Ritchie, K.,
Blanche, H., Dartigues, J.F., Tzourio, C., Gut, I., van Broeckhoven, C.,
Alperovitch, A., Lathrop, M., and Amouyel, P., 2009. Genome-wide association
study identifies variants at CLU and CR1 associated with Alzheimer's disease.
Nat.Genet. 41, 1094-1099.
Laycock, K.A., Brady, R.H., Lee, S.F., Osborne, P.A.,
Johnson, E.M., and Pepose, J.S., 1994. The role of nerve
growth factor in modulating herpes simplex virus reactivation in vivo.
Graefes Arch.Clin.Exp.Ophthalmol. 232, 421-425.
Lee, H.G., Ueda, M., Miyamoto, Y.,
Yoneda, Y., Perry, G., Smith, M.A., and Zhu, X., 2006.
Aberrant localization of importin alpha1 in hippocampal
neurons in Alzheimer disease. Brain Res. 1124, 1-4.
Lemansky, P., Brix, K., and Herzog,
V., 1999. Subcellular distribution,
secretion, and posttranslational modifications of clusterin in thyrocytes.
Exp.Cell Res. 251, 147-155.
Leskov, K.S., Klokov, D.Y., Li, J., Kinsella, T.J.,
and Boothman, D.A., 2003. Synthesis and functional analyses
of nuclear clusterin, a cell death protein. J.Biol Chem. 278,
11590-11600.
Letenneur, L., Peres, K., Fleury,
H., Garrigue, I., Barberger-Gateau, P., Helmer, C., Orgogozo, J.M., Gauthier,
S., and Dartigues, J.F., 2008. Seropositivity to
herpes simplex virus antibodies and risk of Alzheimer's disease: a
population-based cohort study. PLoS.One. 3, e3637.
Leveugle, B., Spik, G., Perl, D.P., Bouras, C.,
Fillit, H.M., and Hof, P.R., 1994. The iron-binding protein lactotransferrin is
present in pathologic lesions in a variety of neurodegenerative disorders: a
comparative immunohistochemical analysis. Brain Res. 650, 20-31.
Levy, E., Sastre, M., Kumar, A.,
Gallo, G., Piccardo, P., Ghetti, B., and Tagliavini, F., 2001.
Codeposition of cystatin C with amyloid-beta protein in the
brain of Alzheimer disease patients. J.Neuropathol.Exp.Neurol. 60,
94-104.
Li, S., Carpenter, D., Hsiang, C.,
Wechsler, S.L., and Jones, C., 2010.
Herpes simplex virus type 1 latency-associated transcript inhibits apoptosis
and promotes neurite sprouting in neuroblastoma cells following serum
starvation by maintaining protein kinase B (AKT) levels. J.Gen.Virol. 91,
858-866.
Li, X., An, W.L., Alafuzoff,
I., Soininen, H., Winblad, B., and Pei, J.J., 2004. Phosphorylated eukaryotic
translation factor 4E is elevated in Alzheimer brain. Neuroreport.
15, 2237-2240.
Liao, L., Cheng, D., Wang, J., Duong, D.M., Losik,
T.G., Gearing, M., Rees, H.D., Lah, J.J., Levey, A.I., and Peng, J., 2004. Proteomic characterization of postmortem amyloid plaques isolated
by laser capture microdissection. J.Biol Chem. 279, 37061-37068.
Love, S., Barber, R., and Wilcock, G.K., 1999.
Increased poly(ADP-ribosyl)ation of nuclear proteins
in Alzheimer's disease. Brain. 122, 247-253.
Lyman, M.G. and Enquist, L.W., 2009. Herpesvirus interactions with the host cytoskeleton.
J.Virol. 83, 2058-2066.
Ma, J.F., Huang, Y., Chen, S.D., and
Halliday, G., 2010. Immunohistochemical evidence for
macroautophagy in neurons and endothelial cells in Alzheimer's disease. Neuropathol.Appl.Neurobiol.
MacGibbon, G.A., Lawlor, P.A.,
Sirimanne, E.S., Walton, M.R., Connor, B., Young, D., Williams, C., Gluckman,
P., Faull, R.L., Hughes, P., and Dragunow, M., 1997.
Bax expression in mammalian neurons undergoing apoptosis, and
in Alzheimer's disease hippocampus. Brain Res. 750, 223-234.
Manabe, T., Katayama, T., Sato, N.,
Gomi, F., Hitomi, J., Yanagita, T., Kudo, T., Honda, A., Mori, Y., Matsuzaki,
S., Imaizumi, K., Mayeda, A., and Tohyama, M., 2003.
Induced HMGA1a expression causes aberrant splicing of Presenilin-2 pre-mRNA in
sporadic Alzheimer's disease. Cell Death.Differ. 10,
698-708.
Mann, D.M., Tinkler, A.M., and
Yates, P.O., 1983. Neurological disease and herpes
simplex virus. An immunohistochemical study. Acta Neuropathol. 60, 24-28.
Marquez-Sterling, N.R., Lo, A.C.,
Sisodia, S.S., and Koo, E.H., 1997. Trafficking
of cell-surface beta-amyloid precursor protein: evidence that a sorting
intermediate participates in synaptic vesicle recycling. J.Neurosci. 17,
140-151.
McComb, R.D., Miller, K.A., and
Carson, S.D., 1991. Tissue factor antigen in senile
plaques of Alzheimer's disease. Am.J.Pathol. 139, 491-494.
McGeer, P.L., Akiyama, H., Itagaki, S., and McGeer,
E.G., 1989. Activation of the classical complement pathway in
brain tissue of Alzheimer patients. Neurosci.Lett. 107, 341-346.
McGeer, P.L., Walker, D.G., Akiyama,
H., Kawamata, T., Guan, A.L., Parker, C.J., Okada, N., and McGeer, E.G., 1991.
Detection of the membrane inhibitor of reactive lysis (CD59)
in diseased neurons of Alzheimer brain. Brain Res. 544, 315-319.
McNamara, J.O., Grigston, J.C., VanDongen, H.M., and
VanDongen, A.M., 2004. Rapid dendritic transport of TGN38, a
putative cargo receptor. Brain Res.Mol.Brain Res. 127, 68-78.
McShea, A., Zelasko, D.A., Gerst, J.L., and Smith,
M.A., 1999. Signal transduction abnormalities in Alzheimer's disease: evidence of a pathogenic stimuli. Brain Res. 815, 237-242.
Meilinger, M., Gschwentner, C.,
Burger, I., Haumer, M., Wahrmann, M., Szollar, L., Nimpf, J., and Huettinger,
M., 1999. Metabolism of activated complement component C3 is
mediated by the low density lipoprotein receptor-related protein/alpha(2)-macroglobulin receptor. J.Biol.Chem. 274,
38091-38096.
Miller, R.M. and Federoff, H.J., 2008.
Isoform-specific effects of ApoE on HSV immediate early gene expression and
establishment of latency. Neurobiol.Aging 29, 71-77.
Mori, I., Kimura, Y., Naiki, H.,
Matsubara, R., Takeuchi, T., Yokochi, T., and Nishiyama, Y., 2004.
Reactivation of HSV-1 in the brain of patients with familial
Alzheimer's disease. J.Med.Virol. 73, 605-611.
Muller, T., Meyer, H.E.,
Egensperger, R., and Marcus, K., 2008.
The amyloid precursor protein intracellular domain (AICD) as
modulator of gene expression, apoptosis, and cytoskeletal dynamics-relevance
for Alzheimer's disease. Prog.Neurobiol. 85, 393-406.
Namba, Y., Tomonaga, M., Kawasaki,
H., Otomo, E., and Ikeda, K., 1991. Apolipoprotein
E immunoreactivity in cerebral amyloid deposits and neurofibrillary tangles in
Alzheimer's disease and kuru plaque amyloid in Creutzfeldt-Jakob disease.
Brain Res. 541, 163-166.
Nikaido, T., Austin, J., Trueb, L.,
Hutchison, J., Rinehart, R., Stuckenbrok, H., and Miles, B., 1970.
Isolation and preliminary characterization of Alzheimer
plaques from presenile and senile dementia. Trans.Am.Neurol.Assoc. 95,
47-50.
Okamoto, K., Hirai, S., Iizuka, T., Yanagisawa, T.,
and Watanabe, M., 1991. Reexamination of granulovacuolar
degeneration. Acta Neuropathol. 82, 340-345.
Padula, M.E., Sydnor, M.L., and
Wilson, D.W., 2009. Isolation and preliminary
characterization of herpes simplex virus 1 primary enveloped virions from the
perinuclear space. J.Virol. 83, 4757-4765.
Page, G., Rioux, B.A., Ingrand, S., Lafay-Chebassier,
C., Pain, S., Perault Pochat, M.C., Bouras, C., Bayer, T., and Hugon, J., 2006.
Activated double-stranded RNA-dependent protein kinase and
neuronal death in models of Alzheimer's disease. Neuroscience.
139, 1343-1354.
Pangalos, M.N., Shioi, J., Efthimiopoulos, S., Wu, A.,
and Robakis, N.K., 1996. Characterization of appican, the
chondroitin sulfate proteoglycan form of the Alzheimer amyloid precursor
protein. Neurodegeneration. 5, 445-451.
Paul, S., 2008.
Dysfunction of the ubiquitin-proteasome system in multiple disease conditions:
therapeutic approaches. Bioessays. 30, 1172-1184.
Peel, A.L. and Bredesen, D.E., 2003. Activation of the
cell stress kinase PKR in Alzheimer's disease and human amyloid precursor
protein transgenic mice. Neurobiol.Dis. 14, 52-62.
Pei, J.J., Braak, H., Gong, C.X.,
Grundke-Iqbal, I., Iqbal, K., Winblad, B., and Cowburn, R.F., 2002.
Up-regulation of cell division cycle (cdc) 2 kinase in
neurons with early stage Alzheimer's disease neurofibrillary degeneration. Acta Neuropathol. 104, 369-376.
Perl, D.P., 2010. Neuropathology of
Alzheimer's disease. Mt.Sinai J.Med. 77, 32-42.
Perlmutter, L.S., Scott, S.A.,
Barron, E., and Chui, H.C., 1992. MHC class
II-positive microglia in human brain: association with Alzheimer lesions.
J.Neurosci.Res. 33, 549-558.
Perry, G., Friedman, R., Shaw, G.,
and Chau, V., 1987. Ubiquitin is detected in neurofibrillary tangles and
senile plaque neurites of Alzheimer disease brains. Proc.Natl.Acad.Sci.U.S.A.
84, 3033-3036.
Perry, G., Siedlak, S.L., Richey, P., Kawai, M., Cras,
P., Kalaria, R.N., Galloway, P.G., Scardina, J.M., Cordell, B., Greenberg,
B.D., and ., 1991. Association of heparan sulfate
proteoglycan with the neurofibrillary tangles of Alzheimer's disease. J.Neurosci. 11, 3679-3683.
Piacentini, R., Civitelli,
L., Ripoli, C., Elena, M.M., De Chiara, G., Garaci, E., Azzena, G.B., Palamara,
A.T., and Grassi, C., 2010. HSV-1 promotes Ca(2+)-mediated
APP phosphorylation and Abeta accumulation in rat cortical neurons. Neurobiol.Aging.
Powers, J.H., Buster, B.L., Reist,
C.J., Martin, E., Bridges, M., Sutherland, W.M., Taylor, R.P., and Scheld,
W.M., 1995. Complement-independent binding of
microorganisms to primate erythrocytes in vitro by cross-linked monoclonal
antibodies via complement receptor 1. Infect.Immun. 63, 1329-1335.
Pyles, R.B., 2001. The association of herpes simplex
virus and Alzheimer's disease: a potential synthesis of genetic and
environmental factors. Herpes. 8, 64-68.
Rajendran, L., Honsho, M., Zahn,
T.R., Keller, P., Geiger, K.D., Verkade, P., and Simons, K., 2006.
Alzheimer's disease beta-amyloid peptides are released in association with
exosomes. Proc.Natl.Acad.Sci.U.S.A. 103, 11172-11177.
Rogers, J., Li, R., Mastroeni, D.,
Grover, A., Leonard, B., Ahern, G., Cao, P., Kolody, H., Vedders, L., Kolb,
W.P., and Sabbagh, M., 2006. Peripheral
clearance of amyloid beta peptide by complement C3-dependent adherence to
erythrocytes. Neurobiol.Aging 27, 1733-1739.
Ronco, P. and Debiec, H., 2007. Podocyte
antigens and glomerular disease. Nephron Exp.Nephrol.
107, e41-e46.
Saffran, H.A., Pare, J.M., Corcoran, J.A., Weller,
S.K., and Smiley, J.R., 2007. Herpes simplex virus eliminates host
mitochondrial DNA. EMBO Rep. 8, 188-193.
Sahara, N., Maeda, S., Yoshiike, Y., Mizoroki, T.,
Yamashita, S., Murayama, M., Park, J.M., Saito, Y., Murayama, S., and
Takashima, A., 2007. Molecular chaperone-mediated tau protein metabolism
counteracts the formation of granular tau oligomers in human brain. J.Neurosci.Res. 85, 3098-3108.
Saito, Y. and Price, R.W., 1984. Enhanced regional
uptake of 2-deoxy-D-[14C]glucose in focal herpes
simplex type 1 encephalitis: autoradiographic study in the rat. Neurology. 34, 276-284.
Santpere, G., Nieto, M., Puig, B.,
and Ferrer, I., 2006. Abnormal Sp1 transcription factor
expression in Alzheimer disease and tauopathies. Neurosci.Lett. 397,
30-34.
Satpute-Krishnan, P., DeGiorgis,
J.A., and Bearer, E.L., 2003. Fast anterograde
transport of herpes simplex virus: role for the amyloid precursor protein of alzheimer's disease. Aging Cell 2, 305-318.
Savage, M.J., Iqbal, M., Loh, T.,
Trusko, S.P., Scott, R., and Siman, R., 1994.
Cathepsin G: localization in human cerebral cortex and generation of
amyloidogenic fragments from the beta-amyloid precursor protein. Neuroscience. 60, 607-619.
Schillinger, J.A., Xu, F.,
Sternberg, M.R., Armstrong, G.L., Lee, F.K., Nahmias, A.J., McQuillan, G.M.,
Louis, M.E., and Markowitz, L.E., 2004.
National seroprevalence and trends in herpes simplex virus type 1 in the United
States, 1976-1994. Sex Transm.Dis. 31, 753-760.
Schindowski, K., Belarbi, K., and Buee, L., 2008.
Neurotrophic factors in Alzheimer's disease: role of axonal transport. Genes Brain Behav. 7 Suppl 1, 43-56.
Sheffield, L.G. and Mirra, S.S., 2008. The distribution of karyopherin beta proteins in Alzheimer's
disease.� 22 ed.,� p. 707.6.
Shioi, J., Pangalos, M.N., Menelas, N., Wu, A., and
Robakis, N.K., 1996. Structure and Function of Appican, The
Proteoglycan Form of the Alzheimer Amyloid Precursor Protein.� 8 ed.,� pp. 253-263.
Sims, N.R., Bowen, D.M., Smith, C.C., Flack, R.H.,
Davison, A.N., Snowden, J.S., and Neary, D., 1980. Glucose
metabolism and acetylcholine synthesis in relation to neuronal activity in
Alzheimer's disease. Lancet. 1, 333-336.
Singh, T.J., Grundke-Iqbal, I.,
McDonald, B., and Iqbal, K., 1994. Comparison
of the phosphorylation of microtubule-associated protein tau by non-proline
dependent protein kinases. Mol.Cell Biochem. 131, 181-189.
Singhrao, S.K., Neal, J.W., Rushmere, N.K., Morgan,
B.P., and Gasque, P., 1999. Differential expression of
individual complement regulators in the brain and choroid plexus. Lab
Invest. 79, 1247-1259.
Smith, J.S. and Robinson, N.J.,
2002. Age-specific prevalence of infection with herpes
simplex virus types 2 and 1: a global review. J.Infect.Dis.
186 Suppl 1, S3-28.
Snow, A.D., Mar, H., Nochlin, D., Kresse, H., and
Wight, T.N., 1992. Peripheral distribution of dermatan sulfate proteoglycans
(decorin) in amyloid-containing plaques and their presence in neurofibrillary
tangles of Alzheimer's disease. J.Histochem.Cytochem. 40, 105-113.
Sohn, J.H., So, J.O., Hong, H.J.,
Kim, J.W., Na, D.R., Kim, M., Kim, H., Nam, E., Ha, H.J., Kim, Y.H., and
Mook-Jung, I., 2009. Identification of autoantibody
against beta-amyloid peptide in the serum of elderly. Front Biosci. 14,
3879-3883.
Stelzl, U., Worm, U., Lalowski, M., Haenig, C.,
Brembeck, F.H., Goehler, H., Stroedicke, M., Zenkner, M., Schoenherr, A.,
Koeppen, S., Timm, J., Mintzlaff, S., Abraham, C., Bock, N., Kietzmann, S.,
Goedde, A., Toksoz, E., Droege, A., Krobitsch, S., Korn, B., Birchmeier, W.,
Lehrach, H., and Wanker, E.E., 2005. A human protein-protein interaction
network: a resource for annotating the proteome. Cell 122, 957-968.
Stenmark, H., 2009. Rab GTPases as
coordinators of vesicle traffic. Nat.Rev.Mol.Cell Biol. 10, 513-525.
Su, J.H., Zhao, M., Anderson, A.J.,
Srinivasan, A., and Cotman, C.W., 2001.
Activated caspase-3 expression in Alzheimer's and aged
control brain: correlation with Alzheimer pathology. Brain Res. 898,
350-357.
Suenaga, T., Satoh, T., Somboonthum,
P., Kawaguchi, Y., Mori, Y., and Arase, H., 2010.
Myelin-associated glycoprotein mediates membrane fusion and entry of
neurotropic herpesviruses. Proc.Natl.Acad.Sci.U.S.A 107, 866-871.
Szodorai, A., Kuan, Y.H., Hunzelmann, S., Engel, U.,
Sakane, A., Sasaki, T., Takai, Y., Kirsch, J., Muller, U., Beyreuther, K.,
Brady, S., Morfini, G., and Kins, S., 2009. APP anterograde transport requires
Rab3A GTPase activity for assembly of the transport vesicle. J.Neurosci. 29,
14534-14544.
Takami, K., Matsuo, A., Terai, K.,
Walker, D.G., McGeer, E.G., and McGeer, P.L., 1998.
Fibroblast growth factor receptor-1 expression in the cortex
and hippocampus in Alzheimer's disease. Brain Res. 802, 89-97.
Takata, K., Kitamura, Y., Kakimura,
J., Shibagaki, K., Tsuchiya, D., Taniguchi, T., Smith, M.A., Perry, G., and
Shimohama, S., 2003. Role of high mobility group
protein-1 (HMG1) in amyloid-beta homeostasis.
Biochem.Biophys.Res.Commun. 301, 699-703.
Tan, Z., Sun, N., and Schreiber, S.S., 1998. Immunohistochemical localization of redox factor-1 (Ref-1) in
Alzheimer's hippocampus. Neuroreport. 9,
2749-2752.
Tanimukai, H., Grundke-Iqbal, I.,
and Iqbal, K., 2005. Up-regulation of inhibitors of
protein phosphatase-2A in Alzheimer's disease. Am.J.Pathol. 166,
1761-1771.
Tebar, F., Bohlander, S.K., and
Sorkin, A., 1999. Clathrin assembly lymphoid myeloid leukemia (CALM)
protein: localization in endocytic-coated pits, interactions with clathrin, and
the impact of overexpression on clathrin-mediated traffic. Mol.Biol.Cell
10, 2687-2702.
Thal, D.R., Schober, R., and Birkenmeier, G., 1997. The subunits of alpha2-macroglobulin receptor/low density
lipoprotein receptor-related protein, native and transformed
alpha2-macroglobulin and interleukin 6 in Alzheimer's disease. Brain
Res. 777, 223-227.
Thompson, P.M., Hayashi, K.M., de Zubicaray, G.,
Janke, A.L., Rose, S.E., Semple, J., Herman, D., Hong, M.S., Dittmer, S.S.,
Doddrell, D.M., and Toga, A.W., 2003. Dynamics of gray matter
loss in Alzheimer's disease. J.Neurosci. 23, 994-1005.
Ueberham, U., Ueberham, E.,
Gruschka, H., and Arendt, T., 2006. Altered
subcellular location of phosphorylated Smads in Alzheimer's disease. Eur.J.Neurosci. 24, 2327-2334.
Umahara, T., Uchihara, T., Tsuchiya,
K., Nakamura, A., Iwamoto, T., Ikeda, K., and Takasaki, M., 2004.
14-3-3 proteins and zeta isoform containing neurofibrillary
tangles in patients with Alzheimer's disease. Acta
Neuropathol. 108, 279-286.
van
der Wal, E.A., Gomez-Pinilla, F., and Cotman, C.W., 1993. Transforming growth
factor-beta 1 is in plaques in Alzheimer and Down
pathologies. Neuroreport. 4, 69-72.
Van Hoesen, G.W. and Hyman, B.T., 1990. Hippocampal
formation: anatomy and the patterns of pathology in Alzheimer's disease.
Prog.Brain Res. 83, 445-457.
Vecchi, M., Polo, S., Poupon, V., van de Loo, J.W.,
Benmerah, A., and Di Fiore, P.P., 2001. Nucleocytoplasmic
shuttling of endocytic proteins. J.Cell Biol. 153, 1511-1517.
Veerhuis, R., Janssen, I., De Groot, C.J., Van
Muiswinkel, F.L., Hack, C.E., and Eikelenboom, P., 1999. Cytokines associated
with amyloid plaques in Alzheimer's disease brain stimulate human glial and
neuronal cell cultures to secrete early complement proteins, but not
C1-inhibitor. Exp.Neurol. 160, 289-299.
Verbeek, M.M., Otte-Holler, I., van den, B.J., van den
Heuvel, L.P., David, G., Wesseling, P., and de Waal, R.M., 1999. Agrin is a
major heparan sulfate proteoglycan accumulating in Alzheimer's disease brain. Am.J.Pathol. 155, 2115-2125.
Vicetti Miguel, R.D., Sheridan,
B.S., Harvey, S.A., Schreiner, R.S., Hendricks, R.L., and Cherpes, T.L., 2010.
17-beta estradiol promotion of herpes simplex virus type 1 reactivation is
estrogen receptor dependent. J.Virol. 84, 565-572.
Vidal, M., Goudreau, N., Cornille,
F., Cussac, D., Gincel, E., and Garbay, C., 1999.
Molecular and cellular analysis of Grb2 SH3 domain mutants: interaction with Sos and dynamin. J.Mol.Biol. 290, 717-730.
Vukic, V., Callaghan, D., Walker,
D., Lue, L.F., Liu, Q.Y., Couraud, P.O., Romero, I.A., Weksler, B.,
Stanimirovic, D.B., and Zhang, W., 2009.
Expression of inflammatory genes induced by beta-amyloid peptides in human
brain endothelial cells and in Alzheimer's brain is mediated by the JNK-AP1
signaling pathway. Neurobiol.Dis. 34, 95-106.
Wang, J. and Maldonado, M.A., 2006. The
ubiquitin-proteasome system and its role in inflammatory and autoimmune
diseases. Cell Mol.Immunol. 3, 255-261.
Wang, Q., Woltjer, R.L., Cimino, P.J., Pan, C.,
Montine, K.S., Zhang, J., and Montine, T.J., 2005. Proteomic analysis of
neurofibrillary tangles in Alzheimer disease identifies GAPDH as a
detergent-insoluble paired helical filament tau binding protein. FASEB J. 19,
869-871.
Williams, P., Verhagen, J., and Elliott, G., 2008. Characterization of a CRM1-dependent nuclear export signal in the C
terminus of herpes simplex virus type 1 tegument protein UL47. J.Virol.
82, 10946-10952.
Wojtowicz, W.M., Farzan, M., Joyal,
J.L., Carter, K., Babcock, G.J., Israel, D.I., Sodroski, J., and Mirzabekov,
T., 2002. Stimulation of enveloped virus
infection by beta-amyloid fibrils. J.Biol.Chem. 277, 35019-35024.
Wozniak, M.A., Frost, A.L., and Itzhaki, R.F., 2009.
Alzheimer's disease-specific tau phosphorylation is induced by herpes simplex
virus type 1. J.Alzheimers.Dis. 16, 341-350.
Wozniak, M.A., Itzhaki, R.F., Shipley, S.J., and
Dobson, C.B., 2007. Herpes simplex virus infection causes cellular beta-amyloid
accumulation and secretase upregulation. Neurosci.Lett. 429, 95-100.
Wozniak, M.A., Mee, A.P., and Itzhaki, R.F., 2009.
Herpes simplex virus type 1 DNA is located within Alzheimer's disease amyloid
plaques. J.Pathol. 217, 131-138.
Wozniak, M.A., Shipley, S.J.,
Combrinck, M., Wilcock, G.K., and Itzhaki, R.F., 2005.
Productive herpes simplex virus in brain of elderly normal
subjects and Alzheimer's disease patients. J.Med.Virol. 75, 300-306.
Wu, Q., Hu, H., Lan, J.,
Emenari, C., Wang, Z., Chang, K.S., Huang, H., and Yao, X., 2009. PML3
Orchestrates the Nuclear Dynamics and Function of TIP60. J.Biol.Chem.
284, 8747-8759.
Yamazaki, Y., Takahashi, T., Hiji,
M., Kurashige, T., Izumi, Y., Yamawaki, T., and Matsumoto, M., 2010.
Immunopositivity for ESCRT-III subunit CHMP2B in
granulovacuolar degeneration of neurons in the Alzheimer's disease hippocampus.
Neurosci.Lett.
Yasuhara, O., Walker, D.G., and McGeer, P.L., 1994.
Hageman factor and its binding sites are present in senile plaques of
Alzheimer's disease. Brain Res. 654, 234-240.
Young, A.B., 1987. Cortical amino
acidergic pathways in Alzheimer's disease. J.Neural Transm.Suppl. 24,
147-152.
Zambrano, A., Solis, L., Salvadores,
N., Cortes, M., Lerchundi, R., and Otth, C., 2008.
Neuronal cytoskeletal dynamic modification and neurodegeneration induced by
infection with herpes simplex virus type 1. J.Alzheimers.Dis. 14, 259-269.
Zanjani, H., Finch, C.E., Kemper, C., Atkinson, J.,
McKeel, D., Morris, J.C., and Price, J.L., 2005. Complement activation in very
early Alzheimer disease. Alzheimer Dis.Assoc.Disord. 19, 55-66.
Zheng, P., Eastman, J., Vande, P.S.,
and Pimplikar, S.W., 1998. PAT1, a microtubule-interacting protein, recognizes
the basolateral sorting signal of amyloid precursor protein.
Proc.Natl.Acad.Sci.U.S.A 95, 14745-14750.
View Stats
